 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_23792-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1475aa MW: 163237 Da PI: 7.0947 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178.2 | 1e-55 | 21 | 136 | 3 | 117 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
+e ++rwl+++ei++iL n++k++++ e+++rp+sgsl+L++rk++ryfrkDG+ w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fq
TRIUR3_23792-P1 21 QEaQNRWLRPTEICQILSNYKKFSIAPEPPNRPPSGSLFLFDRKILRYFRKDGHIWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQ 114
5559****************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylev 117
rr+ywlLee + +ivlvhyle
TRIUR3_23792-P1 115 RRTYWLLEEGFMNIVLVHYLEI 136
*******************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.103 | 16 | 142 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.2E-77 | 19 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.5E-50 | 22 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.5E-7 | 444 | 531 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 4.4E-7 | 445 | 518 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 8.56E-18 | 445 | 530 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 2.8E-16 | 622 | 737 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.72E-12 | 623 | 735 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 1.3E-16 | 632 | 738 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.396 | 643 | 737 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.458 | 676 | 708 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.069 | 676 | 705 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 420 | 715 | 744 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 5.58E-8 | 847 | 900 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 1.5 | 849 | 871 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0083 | 851 | 870 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.627 | 851 | 879 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 872 | 894 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.651 | 873 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 9.8E-5 | 874 | 894 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF50104 | 2.32E-5 | 1079 | 1115 | IPR008991 | Translation protein SH3-like domain |
| Gene3D | G3DSA:2.30.30.30 | 1.0E-5 | 1079 | 1113 | IPR014722 | Ribosomal protein L2 domain 2 |
| Gene3D | G3DSA:3.20.20.100 | 9.2E-7 | 1417 | 1474 | IPR023210 | NADP-dependent oxidoreductase domain |
| SuperFamily | SSF51430 | 5.11E-6 | 1424 | 1474 | IPR023210 | NADP-dependent oxidoreductase domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1475 aa Download sequence Send to blast |
MAEMHKYGLS NQPPDIPQIL QEAQNRWLRP TEICQILSNY KKFSIAPEPP NRPPSGSLFL 60 FDRKILRYFR KDGHIWRKKK DGKTVKEAHE KLKVGSVDVL HCYYAHGEEN ENFQRRTYWL 120 LEEGFMNIVL VHYLEIKGGK QSFSRSKEAE DSPACSNSFA SQSQVASQTM DAESPYSGQI 180 SEYEDAETGY QDENQASTAN ISNNFVTHHG IASVFNDVGA GLRSGSKTAL DSVHFGEPFP 240 EYPTGFTEPT LYSSVTTMGS NNLDDNSRLE TLMTEALYTN NLTQKETDAL SAAGMTSSQR 300 RPPEAPVHSG RLKRADNVKR GLKDWNITKE LAMDRGAWKS TCQNHELVHN DSYTDGSMGY 360 PLLKQSSLDL FKIEPNGLKK FDSFSKWMSD ELAADLDIKS TSDAFWSSTE TVNVADGSSM 420 SIPMNEQLDA YVVSPSLSQD QLFSIIDVSP SWAYTGSQNK VLITGTFLTN KEHVENCKWS 480 CMFGDVEVPV EVLADGSLRC YTPVHQSGRV PFYVTCSNRV ACSEVREFEF HDSETQHMEA 540 ADPHITGINE MHLHIRLEKL LSLGPDDYEK YVMSGGNEKS ELISTIGSLM LDDKFTNLSA 600 PSDEEFSAAQ DKNLEKSVKD KLYYWLIHKI HDDGKGPNVL GKEGQGVIHL VAALGYDWAI 660 RPIIAAGVHV NFRDVRGWTA LHWAASCGRE RTVGALITNG AAAGALTDPT PHFLSGRTPA 720 DLASDNGHKG IAGFLAESAL TSHLSALTLK EAKGCNVEEI CGSIEADGFA EPSSAQLSRQ 780 DSQAESLKDS LSAVRKSTLA ASKIFQAFRV ESFHRKKVVE YGDDDCGLSD ERTLSLVSLK 840 NTKSGQNDMP HSAAVRIQNK FRGWKGRKEF MIIRQKIIKI QAHIRGHQVR RNYKKVVWSV 900 GIVEKVILRW RRKGRGLRGF QPDKQLEGPS SQIQPAEGGS AEGEDEYDFL KDGRKQAEGR 960 LQRSLARVKS MTQYPEAREQ YSRLQACVTE LQESKTLSSS PRSLTCSFDS EQQDEDDDTP 1020 HMVELDGRRC LCRCGLAGVL HVVYDGDLLA RAFSVAFSLV VLVVDHGCLV FVLVQPFKRF 1080 VLIGWVALVN YGKDYGRLVV IVDVVDQNRF RISSLRVPPL AVDLSCSSRI ALLYTYTTVS 1140 WYIPKNVEDT VHREVEIMQH LSGHPAVVTL KAVCGGCKSK LPLCAHVVLA LVCVVEWCLK 1200 GTRRLGVRDY ISLLSLIFCP KYKCINLIMG FSVMAAKRQH GVEMQVVSTL KALRSVRALG 1260 SMQNIGDVAV VLARLIRSNL GPAPAMYGAL DAVLSLCNCN SMTLLRLFCG VRGDTSDQAA 1320 AAYSLGDLHK GAAKISNDIL DSVGVSVYAT TKNSTTVYAK NGAICVCCNL LILTRVECPT 1380 FPKMAPETCP QPEAAKLPYK KLGDSDLLIS EITLGTLQFN QMEAKRPMCG SMESFQELIG 1440 KGKVQYIGVS NETTYGVMQF VHAAKLDGLP KIMSI |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
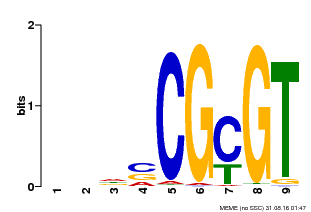
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020196708.1 | 0.0 | calmodulin-binding transcription activator 2-like isoform X1 | ||||
| TrEMBL | M7ZWK4 | 0.0 | M7ZWK4_TRIUA; Calmodulin-binding transcription activator 2 | ||||
| STRING | TRIUR3_23792-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||



