 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_20609-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 303aa MW: 33287.3 Da PI: 6.8997 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53 | 8.1e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +lv +++++G+g+W++++ g+ R+ k+c++rw +yl
TRIUR3_20609-P1 14 KGPWTPEEDIILVSYIQEHGPGNWRSVPVSTGLMRCSKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 44.7 | 3.1e-14 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T E+ +v++ +lG++ W++Ia++++ +Rt++++k++w+++l
TRIUR3_20609-P1 67 RGNFTSHEEGVIVHLQSLLGNR-WAAIASYLP-RRTDNDIKNYWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-22 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 22.093 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 8.59E-29 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.1E-12 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.3E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.03E-11 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.9E-22 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.4E-12 | 66 | 114 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 18.363 | 66 | 116 | IPR017930 | Myb domain |
| Pfam | PF00249 | 9.6E-12 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.59E-9 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0010468 | Biological Process | regulation of gene expression | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 303 aa Download sequence Send to blast |
MGRPPCCDKL GVKKGPWTPE EDIILVSYIQ EHGPGNWRSV PVSTGLMRCS KSCRLRWTNY 60 LRTGIKRGNF TSHEEGVIVH LQSLLGNRWA AIASYLPRRT DNDIKNYWNT HLKKKLRKQQ 120 AMGAIFAPPA SSSPGTAGDN FDHHNHHDMV SRADGYGAGQ AYMNTEVSQL IAGRGHSPFA 180 DAAPEACSSS SYASSVDNIS KLLGGFMKSS TPPPPPHNNY DADDVKPLLP LDSMSGTGSA 240 ELSFTASVQQ PALMGGRVGY EYDDETKQKQ QHQAPLSSIE KWLFDEAADL ELSDECYSVP 300 MLF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 1e-23 | 14 | 116 | 4 | 105 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the regulation of gene (e.g. drought-regulated and flavonoid biosynthetic genes) expression and stomatal movements leading to negative regulation of responses to drought and responses to other physiological stimuli (e.g. light). {ECO:0000269|PubMed:22018045}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00579 | DAP | Transfer from AT5G62470 | Download |
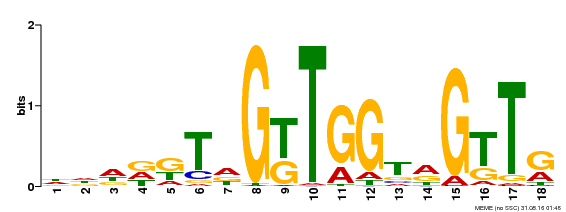
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by abscisic acid (ABA) and osmotic stress (salt stress). {ECO:0000269|PubMed:22018045}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK356090 | 0.0 | AK356090.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1029P14. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020187926.1 | 0.0 | transcription factor MYB30-like | ||||
| Swissprot | B3VTV7 | 5e-88 | MYB60_VITVI; Transcription factor MYB60 | ||||
| TrEMBL | T1MRR7 | 0.0 | T1MRR7_TRIUA; Uncharacterized protein | ||||
| STRING | TRIUR3_20609-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP229 | 38 | 296 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G08810.1 | 1e-79 | myb domain protein 60 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



