 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_16862-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 317aa MW: 36210.4 Da PI: 10.2951 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.3 | 1.6e-17 | 90 | 137 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT++Ed +lv+ v+++G ++W+ Ia+ g++Rt+k+c++rw +yl
TRIUR3_16862-P1 90 KGPWTEQEDMQLVCTVHLFGERRWDFIAKVSGLNRTGKSCRLRWVNYL 137
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 55.3 | 1.6e-17 | 143 | 188 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgr+T+ E+ l++++++++G++ W++Iar+++ gRt++++k++w++++
TRIUR3_16862-P1 143 RGRMTPHEERLILELHARWGNR-WSRIARKLP-GRTDNEIKNYWRTHM 188
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 18.275 | 85 | 137 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.7E-15 | 89 | 139 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.72E-29 | 89 | 184 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 2.0E-16 | 90 | 137 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.8E-20 | 91 | 144 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.55E-11 | 92 | 137 | No hit | No description |
| PROSITE profile | PS51294 | 25.955 | 138 | 192 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.4E-16 | 142 | 190 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.2E-15 | 143 | 187 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.0E-24 | 145 | 191 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.85E-12 | 145 | 188 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MADKKGGMVS SVPMIMADKK ANTAFHIKRT EKENQGNQIE TSPRFPFPIT SKRLEFIKDL 60 VLKLEPTSTQ RKKSKKTGSQ PIQVREETRK GPWTEQEDMQ LVCTVHLFGE RRWDFIAKVS 120 GLNRTGKSCR LRWVNYLHPG LKRGRMTPHE ERLILELHAR WGNRWSRIAR KLPGRTDNEI 180 KNYWRTHMRK KAQERKRNVS PSPSSSSVTY QSIQPQTPSI MGIGEQELHG GSSCITSILK 240 GTPADMDGYL MDQIWMEIEA PSGVNFHDGK DNSYSSPSGP LLPSPMWDYY SPEAGWKMDE 300 IKMAPQVSYS KGIGPSY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 1e-27 | 87 | 192 | 1 | 105 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00600 | PBM | Transfer from AT5G59780 | Download |
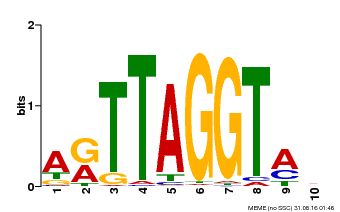
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT009536 | 0.0 | BT009536.1 Triticum aestivum clone wr1.pk0139.g11:fis, full insert mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020201687.1 | 1e-175 | myb-related protein MYBAS2-like isoform X1 | ||||
| Swissprot | Q4JL76 | 1e-128 | MYBA2_ORYSJ; Myb-related protein MYBAS2 | ||||
| TrEMBL | M7ZU89 | 0.0 | M7ZU89_TRIUA; Myb-related protein MYBAS2 | ||||
| STRING | TRIUR3_16862-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP636 | 37 | 169 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G59780.3 | 3e-73 | myb domain protein 59 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



