 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_12807-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 353aa MW: 38864.7 Da PI: 8.4227 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 54.1 | 2.6e-17 | 92 | 145 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rqV WFqNrRa++k
TRIUR3_12807-P1 92 KKRRLNVEQVRTLEKNFEVANKLEPERKMQLARALGLQPRQVALWFQNRRARWK 145
4568999**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 124.8 | 4e-40 | 91 | 180 | 1 | 90 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLree 90
ekkrrl+ eqv++LE++Fe +kLeperK++lar+Lglqprqva+WFqnrRAR+ktkqlEkdy++Lkr++da+k+en++L +++++L++e
TRIUR3_12807-P1 91 EKKRRLNVEQVRTLEKNFEVANKLEPERKMQLARALGLQPRQVALWFQNRRARWKTKQLEKDYDVLKRQFDAVKAENDALLSHNKKLQSE 180
69*************************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 6.5E-19 | 74 | 148 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.682 | 87 | 147 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.59E-18 | 89 | 149 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 3.5E-15 | 90 | 151 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.2E-14 | 92 | 145 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 3.66E-14 | 92 | 148 | No hit | No description |
| PRINTS | PR00031 | 2.2E-5 | 118 | 127 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 122 | 145 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 2.2E-5 | 127 | 143 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.9E-12 | 147 | 180 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 353 aa Download sequence Send to blast |
MAWRPLPPPS SLRISSSTCS KHRRSPFLPS PQCPSLQDFR GGLSPMLGKR PAMYGGGGDG 60 GCGGDEVTGG GGGGGANEEE TSDDGSQLGG EKKRRLNVEQ VRTLEKNFEV ANKLEPERKM 120 QLARALGLQP RQVALWFQNR RARWKTKQLE KDYDVLKRQF DAVKAENDAL LSHNKKLQSE 180 CLQKGCWKRR PCPRLSIYTC EIVIVELSRH SSFSERPSRI WQRPQQILGL KGCREAASEL 240 INLNKETEAS CSNRSENSSE INLDISRTPP SDGPMDAPPS HQQGGGSGGG MIPFYPSVAR 300 PSAGVDIDHL LHASVPKMEH HHGGPDTASF GNLLCGVDEP PPFWPWADHQ QFN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 139 | 147 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
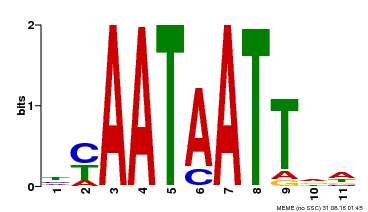
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ353856 | 0.0 | DQ353856.1 Triticum aestivum homeodomain-leucine zipper transcription factor TaHDZipI-2 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020188842.1 | 1e-172 | homeobox-leucine zipper protein HOX21-like isoform X2 | ||||
| Swissprot | Q8S7W9 | 1e-118 | HOX21_ORYSJ; Homeobox-leucine zipper protein HOX21 | ||||
| TrEMBL | M7ZDS3 | 0.0 | M7ZDS3_TRIUA; Homeobox-leucine zipper protein HOX21 | ||||
| STRING | TRIUR3_12807-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2289 | 38 | 94 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 6e-70 | HD-ZIP family protein | ||||



