 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_11462-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 1212aa MW: 131854 Da PI: 6.9655 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 49.7 | 7.5e-16 | 501 | 539 | 2 | 40 |
TCR 2 ekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
++k+C+CkkskClk+YCeCfaag++Cse C+C++C Nk
TRIUR3_11462-P1 501 SCKRCSCKKSKCLKLYCECFAAGVYCSEPCSCQGCLNKP 539
689**********************************96 PP
| |||||||
| 2 | TCR | 48.5 | 1.7e-15 | 586 | 625 | 1 | 40 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
++k+gCnCkks+ClkkYCeC++ g+ Cs++C+Ce CkN+
TRIUR3_11462-P1 586 RHKRGCNCKKSSCLKKYCECYQGGVGCSNNCRCETCKNTF 625
589***********************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 6.6E-17 | 500 | 541 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 37.145 | 501 | 626 | IPR005172 | CRC domain |
| Pfam | PF03638 | 1.5E-11 | 503 | 538 | IPR005172 | CRC domain |
| SMART | SM01114 | 1.6E-15 | 586 | 627 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 4.0E-11 | 588 | 624 | IPR005172 | CRC domain |
| Gene3D | G3DSA:3.40.1280.10 | 8.5E-25 | 990 | 1129 | IPR029026 | tRNA (guanine-N1-)-methyltransferase, N-terminal |
| SuperFamily | SSF75217 | 2.88E-16 | 990 | 1128 | IPR029028 | Alpha/beta knot methyltransferases |
| Pfam | PF00588 | 4.0E-15 | 995 | 1129 | IPR001537 | tRNA/rRNA methyltransferase, SpoU type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001510 | Biological Process | RNA methylation | ||||
| GO:0006396 | Biological Process | RNA processing | ||||
| GO:0009934 | Biological Process | regulation of meristem structural organization | ||||
| GO:0048444 | Biological Process | floral organ morphogenesis | ||||
| GO:0051302 | Biological Process | regulation of cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003723 | Molecular Function | RNA binding | ||||
| GO:0008173 | Molecular Function | RNA methyltransferase activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1212 aa Download sequence Send to blast |
MRVRFALDLL IHLVLETSAP SCSCSVVSAV KIHAYPPAAG RDSPLFSFID SLSPIEPLKS 60 AYSGNGLQAY HQSLNVTSVS SIFTSPHHNA HKESKLSKGS FADYTENELC MEEGTDKNKS 120 PTSSTAVRLF ACTSTITRES HTMTTCSVNE GIVDPPKGPN DLPQPGRFDS GSPDHNTAPC 180 HGVSVRSDLK QDKCPKLEAV QTTNNTVEKR KCLFSTDMQL QDGCQPAKEN NEVMGCEWDD 240 LVSVTSGELL AFDSSMDQHH TGVQLAVNNA ESCGYLLSKL AGGADISDRT HPTTSSQAYY 300 HEMVVGEDKT ENGQLFPEDK KTILSEEIQD NINEENACIP LGCKVETQQR GVRRRCLVFE 360 AAGYSHRTVQ KESAGDLSFS TCKGKSSAQN HRNPGKTPSP HVFRGIGLHL NALALTSKDK 420 MACQDPLATA LVPSLKTEQD VHGNLLSAGG NFFHSGSGSL DLQMDNDDCS VGGFLGNDHN 480 SSQSSSPPKK RRKSDNGDDD SCKRCSCKKS KCLKLYCECF AAGVYCSEPC SCQGCLNKPI 540 HEEIVLSTRK QIEFRNPLAF APKVIRMSEA GQETQEDPKN TPASARHKRG CNCKKSSCLK 600 KYCECYQGGV GCSNNCRCET CKNTFGTRDV AVSAENEEMK QEGEQTESCE KEKENDQQKA 660 NVHSEDHKLV ELVVPITPPL DVSSSLLKQP NFSNAKPPRP CKARSGSSSR SSKASVTVQS 720 RKISKVGDSV FIEEMPDILR EPSSPGIVKT CSPNGKRVSP PHNALSISPN RKGGRKLILK 780 SIPSFPSLVG DTNGGSAICS SDSTTALALV VAEAYLYVCP WNQVHRKSWA GQPIIRDRST 840 TCSPIMKPCE VTNLPEKNEH LIRHDIKMIC VVCGTYLRLA MHLFHDPSSC RKAQDGLYAN 900 GSSQFATREI CRLGFAAIDC LLLLDGAEVP GELHELSGGN VVYVSATVMK KISGMQSVDS 960 TEAIAVMHMP KHFCDLGDDE GGAGLDASFQ SPKRILVLDG IQDPGNLGTL ISNLFKTYVI 1020 SKDGVFLLPA CCDPFNEKAL RAARGASLQL PIISGNWCDL HDLVTRYGMK MLAGHPESSS 1080 DGSEKTHVLS KELADSLMSE SLCLVLGSEG NGLSEETVQA CDLVSIPMEV LLILQYVLVA 1140 RLNQWSATRM SPYETAKTDK MVQTISLILQ SHLCSPSHLQ SHSRPTVAPW NRIHNRRATY 1200 DAWTHDNANV FK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 2e-15 | 503 | 623 | 12 | 120 | Protein lin-54 homolog |
| 5fd3_B | 2e-15 | 503 | 623 | 12 | 120 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 487 | 493 | PKKRRKS |
| 2 | 488 | 492 | KKRRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00624 | PBM | Transfer from PK22848.1 | Download |
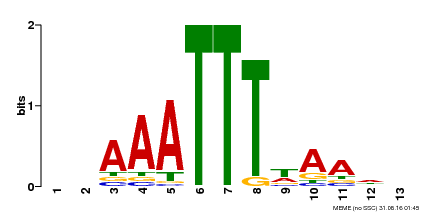
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK375097 | 0.0 | AK375097.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv3085F14. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020167196.1 | 0.0 | protein tesmin/TSO1-like CXC 3 | ||||
| TrEMBL | M7YQ92 | 0.0 | M7YQ92_TRIUA; Protein lin-54-like protein | ||||
| STRING | TRIUR3_11462-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6147 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G14770.1 | 9e-80 | TESMIN/TSO1-like CXC 2 | ||||



