 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_11264-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 486aa MW: 52853.3 Da PI: 7.6482 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 43.9 | 4.8e-14 | 59 | 98 | 2 | 42 |
TCR 2 ekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNkeek 42
+kk+CnCk+s+Clk+YCeCfaag +C+ C+C++C Nk e+
TRIUR3_11264-P1 59 RKKQCNCKNSHCLKLYCECFAAGLYCDG-CNCKQCGNKVEN 98
689*************************.********9875 PP
| |||||||
| 2 | TCR | 50.4 | 4.5e-16 | 141 | 180 | 1 | 40 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
k++kgC+Ckks ClkkYCeCf+a+ Cs++CkC dCkN e
TRIUR3_11264-P1 141 KHNKGCHCKKSGCLKKYCECFQANILCSKNCKCMDCKNFE 180
589***********************************65 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 6.8E-13 | 58 | 98 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 35.49 | 59 | 181 | IPR005172 | CRC domain |
| Pfam | PF03638 | 1.2E-10 | 60 | 95 | IPR005172 | CRC domain |
| SMART | SM01114 | 6.0E-19 | 141 | 182 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 1.4E-12 | 144 | 179 | IPR005172 | CRC domain |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 486 aa Download sequence Send to blast |
MEPEQQHNAQ PPAPPAAAPP QPPQPRPVAP MMHSARSWPL SFTPIKPLVE IKSVTPPKRK 60 KQCNCKNSHC LKLYCECFAA GLYCDGCNCK QCGNKVENEH ARQEAINSTK LRNPKAFQPK 120 IENVPSALSV RKDAGAPSVP KHNKGCHCKK SGCLKKYCEC FQANILCSKN CKCMDCKNFE 180 GSEELQAIIQ GDNASDKNNI QQAANVTLNG AIGSSGYKYS PVRRKRHPED PLGPEANHLD 240 ASQVASSSGL EGCIGYQSRS KMVYRSPLAN TIHPTDVNDL ANHLVIVCRK ATETFLTIAD 300 NKVEMEVERG IYTNADLNND KMKNREVQNG GVSQPDVATH IDQRTAGDLE SPCNNTQEDY 360 RPASPGTQAL LCDEQGTTFG SDYRTSFPSA FHDQDTPELS ALQEKTVLTG FRDYLRLLIT 420 RGKINANNSA GMTEANASSE PAMELDARRN HGATTGPGAS SFPPEAVERP KAPGDPENPR 480 AGDPSA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 1e-26 | 60 | 190 | 11 | 132 | Protein lin-54 homolog |
| 5fd3_B | 1e-26 | 60 | 190 | 11 | 132 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
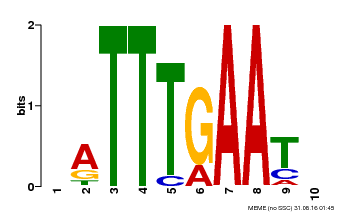
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00045 | PBM | Transfer from AT4G29000 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020150693.1 | 0.0 | protein tesmin/TSO1-like CXC 5 isoform X1 | ||||
| TrEMBL | M7ZEN2 | 0.0 | M7ZEN2_TRIUA; Uncharacterized protein | ||||
| STRING | TRIUR3_11264-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5755 | 26 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G29000.1 | 3e-87 | Tesmin/TSO1-like CXC domain-containing protein | ||||



