 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_09546-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 467aa MW: 51308.6 Da PI: 6.434 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 30.2 | 1e-09 | 182 | 222 | 5 | 45 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL 45
k rr+++NReAAr+sR+RKka+i++Le +L++ ++L
TRIUR3_09546-P1 182 KSLRRLAQNREAARKSRLRKKAYIQNLESSRLKLTQLEQEL 222
788*************************9777777666555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.7E-7 | 178 | 241 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.174 | 180 | 224 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14708 | 3.91E-23 | 182 | 234 | No hit | No description |
| Pfam | PF00170 | 3.5E-7 | 182 | 222 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 3.31E-7 | 182 | 225 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 1.5E-7 | 183 | 226 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 185 | 200 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 4.9E-32 | 264 | 339 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009410 | Biological Process | response to xenobiotic stimulus | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 467 aa Download sequence Send to blast |
MPGFGAPQHT MGLITDLWES VVLCPTDVNI MQPSRVADFG ALAHSAGFRI EDLANFSTNN 60 LFNLKPNTHA YTSEPLQFGN YGKSISPTDL ATTAAAAAAV TAVDPQALLQ QKGAQPNLVA 120 LRTHNNDNWG ESSMADTSPR TDTSTDPDID IDERNQMFEQ GQLAAPTASD SSDKSRDKLD 180 HKSLRRLAQN REAARKSRLR KKAYIQNLES SRLKLTQLEQ ELQRARQQGI FISSSGDQSQ 240 SAGGNGAVAF DMEYARWLEE HNKHINELRA AANAHAGDDD LRKIVDSIMA QYDEFFRLKG 300 VAAKADVFHV LSGMWKTPAE RCFMWLGGFR SSELLKLLAG QLEPLTEQQL TGICNLQQSS 360 QQAEDALSQG MEALQQSLAE TLSSGSLGPA GSSGNVASYM GQMAMAMGKL GTLENFLRQA 420 DNLRLQTLQQ MQRILTTRQS ARALLAISDY FSRLRALSSL WLARPRE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
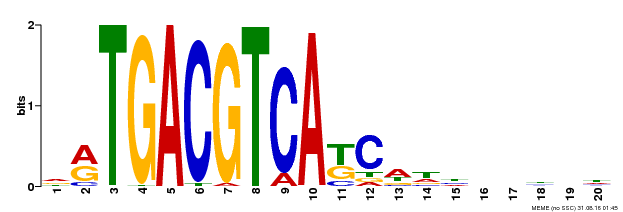
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-TGACG-3'. Recognizes ocs elements like the as-1 motif of the cauliflower mosaic virus 35S promoter. Binding to the as-1-like cis elements mediate auxin- and salicylic acid-inducible transcription. Binds to the hexamer motif 5'-ACGTCA-3' of histone gene promoters. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00492 | DAP | Transfer from AT5G06950 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | WHTHBP1BC1 | 0.0 | D12921.2 Triticum aestivum mRNA for transcription factor HBP-1b(c1), partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020180345.1 | 0.0 | transcription factor HBP-1b(c1) isoform X1 | ||||
| Swissprot | Q41558 | 0.0 | HBP1C_WHEAT; Transcription factor HBP-1b(c1) (Fragment) | ||||
| TrEMBL | M7Z6D7 | 0.0 | M7Z6D7_TRIUA; Transcription factor HBP-1b(C1) | ||||
| STRING | TRIUR3_09546-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP486 | 38 | 185 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06950.4 | 1e-123 | bZIP family protein | ||||



