 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA6619 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 335aa MW: 37028.3 Da PI: 6.6635 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 120.2 | 2.5e-37 | 21 | 137 | 1 | 114 |
TCP 1 aagkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt....ssssaseceaesssss 81
a+g+kdrhsk++T++g+RdRRvRlsa++a++f+d+qd+LG+d++sk+++WL+++ak ai++l ++ + ++++e+ +++ +
Tp57577_TGAC_v2_mRNA6619 21 ATGRKDRHSKVYTAKGPRDRRVRLSAHTAIQFYDVQDRLGYDRPSKAVDWLIKKAKSAIDKLDQLppwnP--IPNATEELQEETN 103
589**************************************************************66431..2233333333333 PP
TCP 82 asnsssg....kaaksaakskksqksaasalnlakes 114
a + ++ + ++++s + +++++++n +++s
Tp57577_TGAC_v2_mRNA6619 104 AVETNEMviaaE---NSESSGYNMQQNNHNHNHHNHS 137
333333354432...3333333333333333333333 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 1.5E-33 | 22 | 136 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 33.172 | 24 | 82 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009793 | Biological Process | embryo development ending in seed dormancy | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 335 aa Download sequence Send to blast |
MGMKSTGGEI VQVQGGHIVR ATGRKDRHSK VYTAKGPRDR RVRLSAHTAI QFYDVQDRLG 60 YDRPSKAVDW LIKKAKSAID KLDQLPPWNP IPNATEELQE ETNAVETNEM VIAAENSESS 120 GYNMQQNNHN HNHHNHSAFI QSSIDNDAIA FFPTSSVAET MNFQSYSSDI ISRTNNSTQD 180 LGLSLHSFQD NSASNDQTLF SESNPVGFDA NFHRIVNWNN DHGTTDMNRT GFMVNSPAFL 240 GQGSAFSAHR GTLQSSFSPS LRPWNDIPMT DSSVDHHHNH KSHSHQPIHQ ASIFGSRFLS 300 DALTGYCIPA RIQGDDESHG VGSERPSPSS NSHH* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 3e-23 | 29 | 82 | 1 | 54 | Putative transcription factor PCF6 |
| 5zkt_B | 3e-23 | 29 | 82 | 1 | 54 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor playing a pivotal role in the control of morphogenesis of shoot organs by negatively regulating the expression of boundary-specific genes such as CUC genes, probably through the induction of miRNA (e.g. miR164) (PubMed:12931144, PubMed:17307931). Required during early steps of embryogenesis (PubMed:15634699). Participates in ovule develpment (PubMed:25378179). Activates LOX2 expression by binding to the 5'-GGACCA-3' motif found in its promoter (PubMed:18816164). {ECO:0000269|PubMed:12931144, ECO:0000269|PubMed:15634699, ECO:0000269|PubMed:17307931, ECO:0000269|PubMed:18816164, ECO:0000269|PubMed:25378179}. | |||||
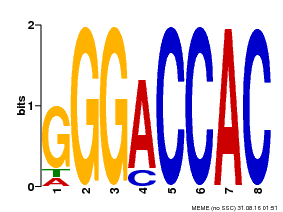
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00062 | PBM | Transfer from AT3G15030 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA6619 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by the miRNA miR-JAW/miR319 (PubMed:12931144, PubMed:18816164). {ECO:0000269|PubMed:12931144, ECO:0000269|PubMed:18816164}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC158498 | 0.0 | AC158498.3 Medicago truncatula chromosome 2 BAC clone mth2-65n13, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013465197.1 | 0.0 | transcription factor TCP4 | ||||
| Swissprot | Q8LPR5 | 9e-71 | TCP4_ARATH; Transcription factor TCP4 | ||||
| TrEMBL | A0A2K3LBT7 | 0.0 | A0A2K3LBT7_TRIPR; Uncharacterized protein | ||||
| STRING | AES83230 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1163 | 34 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G15030.3 | 1e-67 | TCP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA6619 |



