 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA5946 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 505aa MW: 55564.7 Da PI: 7.8914 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 37 | 6.1e-12 | 356 | 402 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ +I +Lq
Tp57577_TGAC_v2_mRNA5946 356 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAITHITDLQ 402
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 4.1E-37 | 51 | 226 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.456 | 352 | 401 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 4.15E-14 | 355 | 406 | No hit | No description |
| SuperFamily | SSF47459 | 3.14E-18 | 355 | 424 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.8E-9 | 356 | 402 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 5.8E-17 | 356 | 422 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.2E-14 | 358 | 407 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 505 aa Download sequence Send to blast |
MASERFWKNE EDKAMLESVL GTEAVAFFNS AVSKHVFSDV IVPPSLDLGI HKRLCHIVKG 60 SKWNYSILWQ VAGLKSGGYV LKYGEGHCQD PKGGPRSEQE RERDDVRRKV LGKIHACWGG 120 SSSIENVYKK LESVSDLYML YLTSVYYVFG FNSQYGPGSS FKCAKPNWSS DAGSCLNQYE 180 SRSFLAKSAG FQTVVFIPLK AGVIELGSME MVPEDMGFLD MVRTTFGEST SGQAKAAPKI 240 FGHELSLGDT KSQSITISFS PKVEDDSGFT SDSYEVQALG ANHAYGNSSN VGVGDSNDAK 300 MFPQLTQMAP GNFTSQARVS SSIDLGNEDS SSPLGDERKP RKRGRKPANG REEPLNHVEA 360 ERQRREKLNQ RFYALRAVVP NISKMDKASL LGDAITHITD LQKKIKLLET EKNMVNNKSQ 420 QLSLQDIDFQ ARQDDAVVRV SCPLDIHPVS GIVKVFREHQ IAAQETNVST AQDKIIHTFS 480 IRTQGGEAAA VQLKEKLEAS LAKN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 3e-29 | 350 | 425 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_B | 3e-29 | 350 | 425 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_E | 3e-29 | 350 | 425 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_F | 3e-29 | 350 | 425 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_G | 3e-29 | 350 | 425 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_I | 3e-29 | 350 | 425 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_M | 3e-29 | 350 | 425 | 2 | 77 | Transcription factor MYC2 |
| 5gnj_N | 3e-29 | 350 | 425 | 2 | 77 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 337 | 345 | RKPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00099 | PBM | Transfer from AT4G16430 | Download |
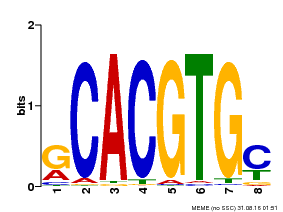
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA5946 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT133685 | 1e-176 | BT133685.1 Medicago truncatula clone JCVI-FLMt-16B12 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013449847.1 | 0.0 | transcription factor bHLH3 | ||||
| Swissprot | A0A3Q7H216 | 1e-171 | MTB3_SOLLC; Transcription factor MTB3 | ||||
| TrEMBL | A0A2K3N6B3 | 0.0 | A0A2K3N6B3_TRIPR; Transcription factor bHLH3-like protein | ||||
| STRING | XP_004493757.1 | 0.0 | (Cicer arietinum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16430.1 | 1e-164 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA5946 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



