 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA5306 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 969aa MW: 110857 Da PI: 7.5592 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 84.8 | 1.4e-26 | 215 | 317 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekd 73
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ +e++ +r+ ++t+Cka+++vk++ d
Tp57577_TGAC_v2_mRNA5306 215 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSfnrprarqhkqeSENSTGRRSCSKTDCKASMHVKRRPD 299
5********************************************999****99877655555558999**************** PP
FAR1 74 gkwevtkleleHnHelap 91
gkw+++++++eHnHel p
Tp57577_TGAC_v2_mRNA5306 300 GKWVIHSFVKEHNHELLP 317
***************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.7E-24 | 215 | 317 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 9.4E-29 | 414 | 506 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.78 | 694 | 730 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.1E-5 | 695 | 728 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 2.1E-7 | 705 | 732 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 969 aa Download sequence Send to blast |
LELGGGRLEL PEEEKQNKKT RKQQNSTPEK KKARKGRTRI GEEENTCADS DKVVQRADCI 60 LVAPPPPLLV HIVKIKVSDT CEINGEITKS EWIGKRNESN HHHHQHNQFF SFNFSVPLKS 120 SKQVLIMDID LRLPSGEHDK EEEETTTLDN MLEGEEKLHN GDIDGRHMVD AGIELHALNG 180 GDLNSPTVDM VMFKEDTNLE PLFGMEFESH GEAYSFYQEY ARSMGFNTAI QNSRRSKTSR 240 EFIDAKFACS RYGTKREYDK SFNRPRARQH KQESENSTGR RSCSKTDCKA SMHVKRRPDG 300 KWVIHSFVKE HNHELLPAQA VSEQTRKMYA VMARQFAEYK TVVGVKNEKN QFDKGRNLGL 360 EFVEAKLMLD FFIQMQSMNS NFFYAVDLGE DQRLKNLLWI DAKSRHDYIN FCDVVSFDTT 420 YVRNKYKMPL VLFVGVNQHY QFILLGCALI SDESAATFSW LLRTWLKGVG GQVPKVIITD 480 HDMTLKSVIS DVIPSACHCV CLWHILGKVS ENLAHVIKKH VNFMTKFEKC VFRSLTSDDF 540 DNRWEKILDR FELRQDECMQ SLYEDRKLWA PTFMKDVFLG GMSTPQRSES VNSFFDKYVH 600 RKTYIQDFVK QYETILQDRY EEEAKADSDT WNKVATLKTP SPLEKSVAGI FTHTVFKKIQ 660 AEIIGAVACH PKLDRQDETT IVHRVHDMET RKDFFVVVNE VNSELSCICR QFEYKGYLCR 720 HALVVLQYSG HSVFPSQYVL KRWTKDAKIR NVTAEESEHM LARVQRYNDL CHRSLKLSEE 780 GSLSQDSYGI AFHALNEAHK NCVSVNNSSK SPTEAGTSGV HGQLSIEEDT QSRNMGKSNK 840 KKNPTKKKKV NSEAEVMTVG ALDNLQQMDK FSARTAVLEG YYGTQQSVQG MLNLMGPTRD 900 DYYGNQQTLQ GLGPINSIPT SHDGYYGAHQ SMPGLAQLDF LRTGFTYGIR DDPNVRAAQL 960 HEDPSRHA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
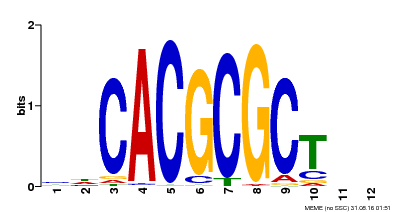
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA5306 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CT030192 | 0.0 | CT030192.6 M.truncatula DNA sequence from clone MTH2-16D16 on chromosome 3, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012571076.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X8 | ||||
| Refseq | XP_027190195.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X4 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A2K3NNZ1 | 0.0 | A0A2K3NNZ1_TRIPR; Protein far-red impaired response 1-like | ||||
| STRING | XP_004499738.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA5306 |



