 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA3133 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1005aa MW: 111709 Da PI: 6.2601 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 127.9 | 3.8e-40 | 157 | 234 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ k+yhrrhkvCe+hska+ +lv + +qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Tp57577_TGAC_v2_mRNA3133 157 VCQVEDCGADLSRGKDYHRRHKVCEMHSKASRALVGNAMQRFCQQCSRFHILQEFDEGKRSCRRRLAGHNKRRRKTNT 234
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.8E-32 | 152 | 219 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.795 | 155 | 232 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.57E-37 | 156 | 236 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 8.9E-29 | 158 | 231 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 7.8E-8 | 754 | 890 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 3.42E-9 | 761 | 889 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.99E-8 | 788 | 889 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1005 aa Download sequence Send to blast |
MEARLGGEGY HFYGVGGSSE LTSMGKRSRE WNLNDWRWDG DLFLASRVNP VPAEGMRVGQ 60 QFFPLGSGIS VAGGSSNTTS SSCSEEGDLE NSKRDKEGEK KRRVIVLEGD GLNEEAGALS 120 LNLAGQASPV VEREIATWDG MNGKKGRVAA GTPNRAVCQV EDCGADLSRG KDYHRRHKVC 180 EMHSKASRAL VGNAMQRFCQ QCSRFHILQE FDEGKRSCRR RLAGHNKRRR KTNTEAVPNG 240 SPTNDDQTSS YLLISLLKIL SNMHSDRSDQ ITDQDLLTHL LRSLASQNDE QGIRNLSNLL 300 REQENLTREG GSSRKSEMVS ALFSNGSQGS PTAITQHQTV SMNRMQQEMM HSHDVTAADH 360 HLISSIKPSI SNSPPASSEA RDSSAPVKMN NFDLNDIYID SDDGTEDLER LPVSTNLGTS 420 SVDYPWAQQD SHQSSPPQTS GNSDSASAQS PSSSSGEAQS RTDRIVFKLF GKEPNEFPLV 480 LKSQILDWLS HSPTDIESYI RPGCIVLTIY LRQAEAVWED LCCDLTSSLN KLLDVSDDPF 540 WRTGWVHIRV QHQMAFIFNG QVVIDTSLPF RSNNYSKIWT VSPIAVPASK RAQFSVKGVN 600 LMRPATRLMC ALEGKYMVCE DAHESTDHYS KELDELQCIQ FSCSVPVTNG RGFIEIEDQG 660 LSSSFFPFIV AEEDVCSEIC VLEPLLELSE TDQDIERTGK IKDKSQAMDF IHEMGWLLHR 720 SQLKSRMVHL NSSVDLIPLE RFTWLMEFSM DHDWCAVVKK LLNLLLDETV NKGDHPTLYQ 780 ALSEMGLLHR AVRKNSKQLV ELLLRYVPDK ALVDGDYQSF LFRPDAVGPA GLTPLHIAAG 840 KDGSEDVLEA LTNDPSMVGI EAWKSARDST GSTPEDYARL RGHYTYIHLV QKKINKRQGA 900 PHVVVEIPST PTESNTNPKQ NESSTSFEIG KAEVRRNQGN CKLCNTKISC RTAVGRSMVY 960 RPAMLSMVAI AAVCVCVALL FKSSPEVLYM FRPFRWESLD FGTS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 5e-30 | 149 | 231 | 2 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 93 | 102 | RDKEGEKKRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
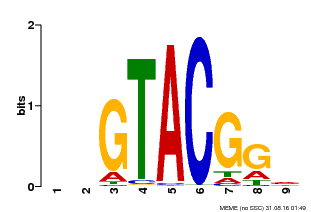
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA3133 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003591325.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A2K3NRP9 | 0.0 | A0A2K3NRP9_TRIPR; Squamosa promoter-binding-like protein 1-like | ||||
| STRING | AES61576 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA3133 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



