 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA28021 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 881aa MW: 100751 Da PI: 6.2188 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 88.4 | 1.1e-27 | 119 | 205 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleH 85
fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e+++ ++r+ ++++t+Cka+++vkk+ dgkw ++++ ++H
Tp57577_TGAC_v2_mRNA28021 119 FYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGVTPESDSG---SSRRPSVKKTDCKACMHVKKKPDGKWTIHEFIKDH 199
****************************************999888...7788889**************************** PP
FAR1 86 nHelap 91
nHel p
Tp57577_TGAC_v2_mRNA28021 200 NHELLP 205
***975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 7.3E-25 | 119 | 205 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 4.1E-29 | 326 | 418 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.285 | 605 | 641 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 5.0E-7 | 607 | 640 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.4E-8 | 616 | 643 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 881 aa Download sequence Send to blast |
LQSGYNFDNF DCGSYSEWND GSVVSGGYPG WNDFDDSVVS GGYPGWNDID DCVVSGGYPG 60 WNDDSAVSGS MGDTVHEEHH TASAPVASPK RDIALFEGDK DFEPPNGIEF ESHEAAYTFY 120 QEYAKSMGFT TSIKNSRRSK KTKEFIDAKF ACSRYGVTPE SDSGSSRRPS VKKTDCKACM 180 HVKKKPDGKW TIHEFIKDHN HELLPALAYH FRIHRNVKLA EKNNIDILHA VSERTRKMYV 240 EMSRQSGGCQ NIGSIVGDMN YQFDRGQFLA LDEGDAQVML EYFKHIQKDN PNFFYSIDLN 300 DEQRLRNLFW IDSKSIDDYL SFNDAVSFDT TYIKSNEKLP FAPFIGVNHH SQPILLGCAL 360 IADETKPTFV WLLKTWLRAM RGKAPKVIVT DQDKALKEAI EEVFPNVRHC FSLWHILEKI 420 PENLSFVIKQ YKNFLPKFNN CIFKSWTDEQ FDLSWWEMVT IFELHDDVWF HSLYEDRKKW 480 VPTYMGDVFL AGMSTPQRSE SMNSFFDKYI HKKITLKEFV KQYGLILQNR YDEEAIADFD 540 TLHKQPALKS PSPWEKQMST VYTHAIFKKF QVEVLGVAGC QSRIEVGDGT VAKFIVQDYE 600 KDEEFLVTWN ELSSEVTCFC RLFEYKGFLC RHALSVLQRC GCSSVPSQYI KKRWTKDAKI 660 REPIITDSTR RVQTRAQRYN DLCKRAIELS EEGSLSEESY NVAIRTLVSC LKNCVLVNNS 720 NDNGAEAGNN GYSLREAEES QSTLALKPSK KRNTTRKRKM QQEQNPVLVD AQDSLQQMDN 780 LTSDAMNLNG YYGTQQNVQG LHVQLNLMEP PQDGYYVNQQ SMQGLGPLNS MGPSHDGFFG 840 TQQSIHGMGG QLEYRPTTTF GYSLQDEHDP HFHSNNSRNT * |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 754 | 759 | TRKRKM |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
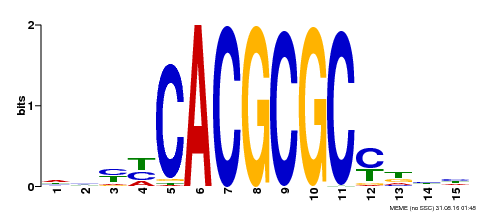
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA28021 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC235177 | 0.0 | AC235177.1 Glycine max strain Williams 82 clone GM_WBb0007H11, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004496891.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A2K3MUT3 | 0.0 | A0A2K3MUT3_TRIPR; Protein far-red impaired response 1-like (Fragment) | ||||
| STRING | XP_004496891.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA28021 |



