 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA17425 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 676aa MW: 73716.5 Da PI: 6.175 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 90.4 | 1.6e-28 | 212 | 265 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Tp57577_TGAC_v2_mRNA17425 212 KPRVVWSVELHQQFVAAVDQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 265
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 76.3 | 1.1e-25 | 34 | 142 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahge 82
vl+vdD+p+ +++l+++l+ y ev+ + +e al ll+e++ +D+++ D+ mp+mdG++ll++i e +lp+i+++a +
Tp57577_TGAC_v2_mRNA17425 34 VLVVDDDPTCLMILEKMLRTCLY-EVTKCNRAETALSLLRENKngFDIVISDVHMPDMDGFKLLEHIGLEM-DLPVIMMSADDG 115
89*********************.***************999999**********************6654.8*********** PP
HHHHHHHHHTTESEEEESS--HHHHHH CS
Response_reg 83 eedalealkaGakdflsKpfdpeelvk 109
++ + + + Ga d+l Kp+ +e+l +
Tp57577_TGAC_v2_mRNA17425 116 KSVVMKGVTHGACDYLIKPVRIEALKN 142
***********************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 4.4E-185 | 14 | 667 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 3.9E-42 | 31 | 161 | No hit | No description |
| SuperFamily | SSF52172 | 3.42E-35 | 31 | 153 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 1.8E-30 | 32 | 144 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.514 | 33 | 148 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 5.4E-23 | 34 | 142 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 3.39E-27 | 35 | 147 | No hit | No description |
| PROSITE profile | PS51294 | 11.518 | 209 | 268 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.63E-20 | 209 | 269 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 5.3E-30 | 211 | 270 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.3E-25 | 212 | 265 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.8E-7 | 214 | 264 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 676 aa Download sequence Send to blast |
MNLSNGKGSI STVTSTATMK SGEGSDQFPA GLRVLVVDDD PTCLMILEKM LRTCLYEVTK 60 CNRAETALSL LRENKNGFDI VISDVHMPDM DGFKLLEHIG LEMDLPVIMM SADDGKSVVM 120 KGVTHGACDY LIKPVRIEAL KNIWQHVVRK KKNEWKDTEQ SGSADEGDRH PKASDDGDYS 180 SSANEGNWRN SRKRRDDEED GDDRDDSSTL KKPRVVWSVE LHQQFVAAVD QLGIDKAVPK 240 KILELMNVPG LTRENVASHL QKYRLYLRRL SGVSQHNNLN NSFLSPQEAS FGTISPMNGL 300 DLQTLAAAGQ LPAQSLATLQ AAGLGRSTVK PGLSMPLMDQ RNLFSFENPR LRFGEGQQQL 360 LSSNKPSNLL HGVPTNMEPK QLANLHQSTQ SLGNLNMRVT GSSTQGNPLL MQMAQSQPRG 420 QMLSEITSPR VPRLPNILAQ PALSNGISDG LLGRNGIASS SRAPSFNPVS QSSSYLNFPM 480 NQSTEMSVSN FPLGSTPGIS SMTNKGSFQP DVNSGIRASA GFPNYDIFND LHQQKSRDWG 540 MASQGPMYDS SSHANPLHGN IDVSPSVLVQ QGYSSSQQPG HNRDPSLIGK SPFSLGEGLD 600 HSNLQNGGQH FNPLIAESST RVKSERVPDA SSQTNLFPDH YGQEDLMSAL LKQEGIVQGE 660 NEFDFDGYSL DNIPV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 3e-23 | 211 | 271 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
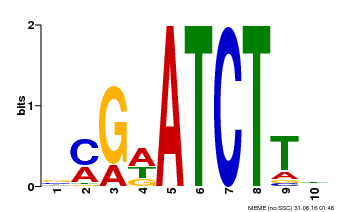
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA17425 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013458296.1 | 0.0 | two-component response regulator ARR2 isoform X2 | ||||
| Swissprot | Q9ZWJ9 | 0.0 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A2Z6N5X6 | 0.0 | A0A2Z6N5X6_TRISU; Two-component response regulator | ||||
| STRING | XP_004508171.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3958 | 34 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 0.0 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA17425 |



