 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA15240 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 520aa MW: 57880.9 Da PI: 7.2153 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 54.9 | 1.6e-17 | 327 | 373 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L++l+P++ K +Ka++Le+A++Y+ksLq
Tp57577_TGAC_v2_mRNA15240 327 VHNLSERRRRDRINEKMKALQQLIPHS-----SKTDKASMLEEAIDYLKSLQ 373
6*************************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.280.10 | 2.4E-19 | 320 | 381 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 3.27E-20 | 321 | 389 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.571 | 323 | 372 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.77E-18 | 326 | 377 | No hit | No description |
| Pfam | PF00010 | 5.1E-15 | 327 | 373 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.5E-19 | 329 | 378 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 520 aa Download sequence Send to blast |
MNNTFPEWNF GSDTFVANQK KQPMGLDHEL VELLWHNGQV VLHSQTNKKP LVNTTSNTSR 60 QVHKNLQSTT LSNLIQADET VSWIQQFPLE DPLEQELCSN LLNELPPCDV ESYNSQTTNT 120 KPSFVEEFSG IPMPAPRFHH HVSDSSQRNN DLCGTNKVVN FSHFSRLPNV SLATSNGDDA 180 KFRDKITGNL SQCDIRECST MTVGSSYCGS NHVQQDPDVS RVSSNGVWTT ISTEPEIIKG 240 DVHKTIPRHE KGKSEMLEPV VTSSSGGSGS SLGKTCSLST RSHGQKRKEI DVEDSVEQSE 300 DTEFKPAVRK KVSQKSGSAR RNRAAEVHNL SERRRRDRIN EKMKALQQLI PHSSKTDKAS 360 MLEEAIDYLK SLQLQLQVMW MGSGMQPMML PGFQQYMSQM SMGMTTPSFP SIQNPMQVPR 420 MPLDPSVSSP QTPNQTLMCQ NPLLGAFNYQ NQIQNPALSE QYARYMGYHL MQNASQPMNM 480 YRYGPQTVQN SQTMIPTSNS SGPMNGAANI NDALSGKMG* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 331 | 336 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
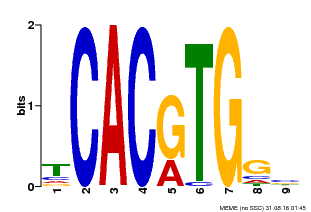
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA15240 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013459610.1 | 0.0 | transcription factor PIF4 isoform X1 | ||||
| TrEMBL | A0A2K3P2D8 | 0.0 | A0A2K3P2D8_TRIPR; Transcription factor PIF5-like protein | ||||
| STRING | XP_004499538.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8146 | 29 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 1e-57 | phytochrome interacting factor 4 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA15240 |



