 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA10803 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1032aa MW: 114685 Da PI: 5.8707 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 141.9 | 1.8e-44 | 5 | 87 | 36 | 118 |
CG-1 36 sgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywlLeeelekivlvhylevk 118
gsl+L++rk++ryfrkDG++w+kkkdgktvrE+hekLKvg+++vl+cyYah+een++fqrr+yw+Le ++ +iv+vhyl+vk
Tp57577_TGAC_v2_mRNA10803 5 GGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVREAHEKLKVGSIDVLHCYYAHGEENENFQRRSYWMLEPDMMHIVFVHYLDVK 87
69******************************************************************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01076 | 5.3E-46 | 1 | 87 | IPR005559 | CG-1 DNA-binding domain |
| PROSITE profile | PS51437 | 63.044 | 1 | 92 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.6E-37 | 5 | 86 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 3.2E-4 | 438 | 526 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.0E-5 | 441 | 515 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 5.29E-16 | 441 | 527 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 7.31E-16 | 629 | 735 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 9.7E-16 | 632 | 736 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.13E-12 | 640 | 772 | No hit | No description |
| PROSITE profile | PS50297 | 17.555 | 641 | 749 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 4.0E-6 | 646 | 734 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.778 | 674 | 706 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.012 | 674 | 703 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 2200 | 713 | 742 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 8.6 | 852 | 874 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 2.92E-7 | 853 | 903 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 8.169 | 854 | 882 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0071 | 855 | 873 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.038 | 875 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.688 | 876 | 900 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.0E-4 | 878 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1032 aa Download sequence Send to blast |
LTISGGSLFL FDRKVLRYFR KDGHNWRKKK DGKTVREAHE KLKVGSIDVL HCYYAHGEEN 60 ENFQRRSYWM LEPDMMHIVF VHYLDVKVNK TNIGASTDTK GVTSDSQNGS SVSSGFPANY 120 GNMPSGSADA MSPTSTLTSL CEDADSEDIH HASSGFHTFR ESQNLGNGPL MDKVDARSNS 180 SYLTHPFSGD HGQLSFSGPN YLPLVQGGKS NQSGATYAEG QRALNIASWD NVMEKSAGLH 240 TDPSLVSSNS IPSSSMGNII EQEHSVFTEG RASQSLHSNW QIPFEDNTGD FPKWSFTQSL 300 NLEFESDYST ELLGKETNNA SSEIGPDLFC FNFEPKEQSV QQNLSKEHTP AQSQDALKSE 360 CGVHGEHSAN YSLNMKHAFM EAESLKKVDS FSRWISKELA AVDDLHMQSS PGISWGTDEC 420 GNVIDDTSLN LSLSQDQLFS IHDFSPKWAY AGSEIEVLII GTFLKSKPEV ATCNWSCMFG 480 EVEVPATVLA NGILCCQAPP HVIGRIPFYV TFSNRFACSE VREFEFREGF TRNVDLADFF 540 NSSTEMTLHL QLEELLTLNS VHLSDQVFED DMEKRNLILK LISLKEEEEY SSNEEPTGDM 600 NISKHRMEVH IFHRQVKEKL YSWLLHKVTE TGKGPHVFGK DGQGVLHLVA ALGYDWAIAP 660 IVTSGVNINF RDVNGCTALH WAASCGRERT VVLLVSMGAA AGALTDPCTA FPSGRTPADI 720 ASSCGHKGIS GFLAESLLTS HLESLTVDDV NKDGAKETLG MKAVQTISER IATPVFSGDI 780 PDPDAICLKD SLDAVRNATQ AADRIHQVYR MQSFQRKQLA QYEGDDEFGL SDQQALSVLA 840 SKASKSGHGD GSVNAAAIQI QKKFRGWTKR KEFLFIRQRV VKIQAHVRGH QVRKKYKSII 900 WSVGILEKVV IRWRRKGSGL RGFRPDAVIK APNQPSNDPV KEDDYDFLKE GRKQSEERFQ 960 KALSRVKSMV QYPEARAQYR RLLNVVDDFR QTKQASTSSP ISSEEAVDGV EDLIDIDMLF 1020 DDDNFLPIVF D* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
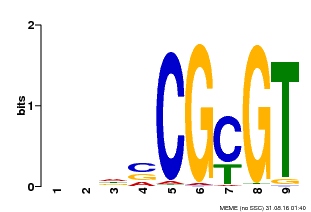
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA10803 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004504800.1 | 0.0 | calmodulin-binding transcription activator 2-like isoform X1 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A2K3NT69 | 0.0 | A0A2K3NT69_TRIPR; Calmodulin-binding transcription activator 2-like protein | ||||
| STRING | XP_004504800.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA10803 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



