 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp7g21990 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 930aa MW: 102529 Da PI: 5.7665 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 91.9 | 4.8e-29 | 564 | 614 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isl++l++yF +++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rkik++
Tp7g21990 564 EKTISLDVLQQYFTGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKIKKV 614
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF55781 | 2.85E-5 | 187 | 258 | IPR029016 | GAF domain-like |
| PROSITE profile | PS51519 | 17.757 | 553 | 634 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 7.0E-26 | 566 | 614 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 3.6E-23 | 831 | 919 | No hit | No description |
| Pfam | PF00564 | 1.7E-20 | 834 | 915 | IPR000270 | PB1 domain |
| SMART | SM00666 | 4.3E-28 | 834 | 916 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 3.1E-27 | 834 | 915 | No hit | No description |
| PROSITE profile | PS51745 | 27.42 | 834 | 916 | IPR000270 | PB1 domain |
| CDD | cd06407 | 6.48E-35 | 835 | 915 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 930 aa Download sequence Send to blast |
MDVDDLDLDG SWPLDQIPYL SSANRMISPI FVSSSSEQPC SPLWAFSDGG GTANHATAGC 60 DDEKISSASG VPSFRLADYP LLLPYSSPSV AENTTEKHNS FQFPSPLMSL VPPENVDNYC 120 VIKERMTQAL RYFKESTEQH VLAQVWAPVR KNGRNLLTTL GQPFVLNPNG NGLNQYRMIS 180 LTYMFSVDSE SDIELGLPGR VFRQKLPEWT PNVQYYSSKE FSRLDHALHY NVRGTLALPV 240 FNPSGESCIG VVELIMTSEK IHYAPEVDRV CKALEAVNLK SSEILDHQTT QICNESRQNA 300 LAEILEVLTV VCETHNLPLA QTWVPCQHGS VLANGGGLKK NCTSFDGSCM GQVCMSTTDM 360 ACYVVDAHVW GFRDACLEHH LQKGQGVAGR AFLNGGSCFC RDITKFCKTQ YPLVHYALMF 420 KLTTCFAISL QSSYTGDDSY ILEFFLPSSI TDDQEQDSLL GSLLVTMKEH FQSLRVASGV 480 DFGEDDDKLS FEIIQALPDK KIHSEIESIR VPFSGFKSSA SETRLIPQLA VQSSDPVNGK 540 INVATVNGVV KEKKKTEKKR GKTEKTISLD VLQQYFTGSL KDAAKSLGVC PTTMKRICRQ 600 HGISRWPSRK IKKVNRSITK LKRVIESVQG TDGGLDLTSM AVSSIPWTHG QTAAQPLNSP 660 SGSKMPELPN TNNSPNHWSS DHSPHEPNGS PELPSNGHKR SRTGDESAGT PTSHGSCDGN 720 QLDETKVPNQ DPLFTVGGSP GLFFPPYSRD HDVSAASFSM PNRLLGTIDH FRGMPIEDAG 780 SSKDLRNLCS SMAFEDKFPE SNWMNNDSNS MYAPPKEEAV ANITREPSGS EMRTVTIKAS 840 YKEDIIRFRI SSGSGIMELK DEVAKRLKLD AVTFDIKYLD DDNEWVLIAC DADLQECLEI 900 PRSSRTNIVR LLVHDLTSNL GSSCESTGEL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
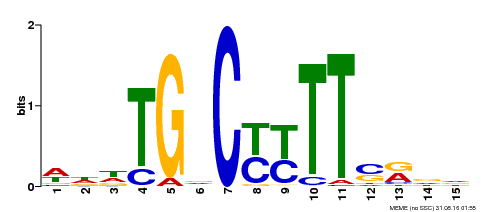
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp7g21990 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK228452 | 0.0 | AK228452.1 Arabidopsis thaliana mRNA for hypothetical protein, complete cds, clone: RAFL15-03-P13. | |||
| GenBank | BT005776 | 0.0 | BT005776.1 Arabidopsis thaliana clone RAFL15-03-P13 (R20174) unknown protein (At4g24020) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006413484.1 | 0.0 | protein NLP7 | ||||
| Refseq | XP_024005488.1 | 0.0 | protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | V4MHY8 | 0.0 | V4MHY8_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006413484.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



