 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp6g33920 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1008aa MW: 113812 Da PI: 6.1923 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 187.8 | 1.1e-58 | 5 | 122 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcyw 100
l+e ++rwl++ ei++iL+n++k+++t+e++trp sgsl+L++rk++ryfrkDG++w+kkkdgkt++E+hekLKvg+++vl+cyYah+e n++fqrrcyw
Tp6g33920 5 LSEaQHRWLRPAEICEILQNYQKFHITSESPTRPASGSLFLFDRKVLRYFRKDGHNWRKKKDGKTIKEAHEKLKVGSIDVLHCYYAHGEGNENFQRRCYW 104
45559*********************************************************************************************** PP
CG-1 101 lLeeelekivlvhylevk 118
+Le++l +iv+vhylevk
Tp6g33920 105 MLEQDLMHIVFVHYLEVK 122
***************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 85.985 | 1 | 127 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 9.7E-87 | 4 | 122 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.8E-51 | 7 | 121 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF01833 | 2.4E-4 | 416 | 496 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 1.92E-14 | 416 | 502 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF12796 | 5.5E-7 | 597 | 675 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 19.863 | 598 | 709 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.3E-17 | 599 | 710 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 4.82E-19 | 599 | 709 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.85E-14 | 610 | 700 | No hit | No description |
| PROSITE profile | PS50088 | 8.683 | 615 | 647 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 2800 | 615 | 644 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 11.327 | 648 | 680 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0018 | 648 | 677 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 3.72E-8 | 822 | 873 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.34 | 823 | 845 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.638 | 825 | 853 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 7.5E-4 | 825 | 844 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0054 | 846 | 868 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.322 | 847 | 871 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.2E-4 | 849 | 868 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1008 aa Download sequence Send to blast |
MEQLLSEAQH RWLRPAEICE ILQNYQKFHI TSESPTRPAS GSLFLFDRKV LRYFRKDGHN 60 WRKKKDGKTI KEAHEKLKVG SIDVLHCYYA HGEGNENFQR RCYWMLEQDL MHIVFVHYLE 120 VKGNRTSIGV KENNSNSVNG TASVNIDSTA SPTSMLSSLC EDADSGDSHQ ASSVFRASPE 180 PQTGNRSGWT SAPGIRIDSH VHGNRDRESD SQRLFDVQAW DGVDNLVTRY DQPCNNLLTQ 240 LQSSNSESML VEERTEKGGL LTAEHLRNPL QTHLNWQIPG QDDLPSPKWP VDLVSHSKMT 300 DDTDLALFEP SAQDHFETFS SLRGSENQLP IGNSFQAPPL SMESEYRPVK KSLLKHEDSL 360 KKVDSFSRWA SKELGEMEDL QLQSSRGDFA WTTVDCETAA DGPSLSPSLS EDQRFTIVDF 420 WPKCAATDAE VKVMVIGTFL LSPQEVTKYN WSCMFGEVEV PAEILVDGVL CCHAPPHTAG 480 QVPFYVTCSN RFACSELREF DFLSETSKKM DVADIYGIYT NESSLQLRIE RLLAQRASIH 540 EHQIFEDVGE KRRKISKIML LKEEKEYLVG MYEKNSTKQE PKERRFREQF EDELYIWLIH 600 KVTEEGKGPN ILDEDGQGIL HFVAALGYDW AIKPILTAGV NINFRDANGW SALHWAAFSG 660 REETVAVLVS LGADARALTD PSPELPLGKT AADLAYVNGH RGISGFLAES SLTSYLEKLT 720 IESKENSSAN SSGPKAVQTV SERTAAPMGY GDVLETLSLK DSLTAVRNAT QAADRLHQVF 780 RMQSFQRKQL SGFGDDDEIG ISNELAVSFA ASKTKNPGHS DVSVHSAAIH IQKKYRGWKK 840 RKEFLIIRQR VVKIQAHVRG HQVRKQYKPI VWSVGLLEKI ILRWRRKGTG LRGFKRNAIP 900 KTVEPEQPVS SLCPTIPQED DYDFLEKGRK QTEERLQKAL TRVKSMVQYP EARDQYRRLL 960 TVVEGFRENE ASSSSSINNR EELVNCEGDD LIDIDSLLND DTLMSISP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds calmodulin in a calcium-dependent manner in vitro (PubMed:11925432). Binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance (PubMed:19270186). Involved in freezing tolerance in association with CAMTA2 and CAMTA3. Contributes together with CAMTA2 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved in drought stress responses by regulating several drought-responsive genes (PubMed:23547968). Involved in auxin signaling and responses to abiotic stresses (PubMed:20383645). Activates the expression of the V-PPase proton pump AVP1 in pollen (PubMed:14581622). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:11925432, ECO:0000269|PubMed:14581622, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:23547968, ECO:0000269|PubMed:23581962, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00501 | DAP | Transfer from AT5G09410 | Download |
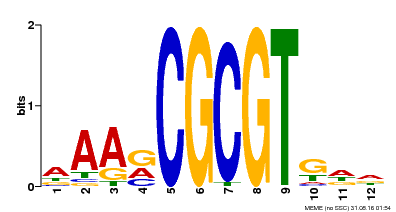
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp6g33920 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by UVB, wounding, ethylene and methyl jasmonate (PubMed:12218065). Induced by salt stress and heat shock (PubMed:12218065, PubMed:20383645). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF491304 | 0.0 | AF491304.1 Brassica napus calmodulin-binding transcription activator (CAMTA) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006399418.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X1 | ||||
| Swissprot | Q9FY74 | 0.0 | CMTA1_ARATH; Calmodulin-binding transcription activator 1 | ||||
| TrEMBL | V4N5T0 | 0.0 | V4N5T0_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006399418.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.2 | 0.0 | ethylene induced calmodulin binding protein | ||||



