 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp5g13610 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1382aa MW: 155011 Da PI: 8.1707 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.2 | 0.0012 | 1265 | 1290 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sFs+ +L H r+ +
Tp5g13610 1265 YQCDmeGCTMSFSSEKQLSLHKRNiC 1290
99********************9877 PP
| |||||||
| 2 | zf-C2H2 | 14.5 | 0.00011 | 1290 | 1312 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk+F ++ +L++H r+H
Tp5g13610 1290 CPvkGCGKTFFSHKYLVQHRRVH 1312
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11 | 0.0013 | 1348 | 1374 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
Tp5g13610 1348 YVCAepSCGQTFRFVSDFSRHKRKtgH 1374
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 2.5E-16 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.952 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 7.9E-14 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 2.1E-47 | 199 | 375 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.633 | 202 | 375 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 2.53E-24 | 217 | 393 | No hit | No description |
| Pfam | PF02373 | 4.6E-34 | 232 | 342 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 6.6E-4 | 1260 | 1287 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 9.4 | 1265 | 1287 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.131 | 1288 | 1317 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0024 | 1288 | 1312 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-6 | 1289 | 1311 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1290 | 1312 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 6.0E-10 | 1304 | 1346 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 4.7E-10 | 1312 | 1339 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0015 | 1318 | 1342 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.928 | 1318 | 1347 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1320 | 1342 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 8.24E-8 | 1336 | 1370 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-8 | 1340 | 1371 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.281 | 1348 | 1379 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.5 | 1348 | 1374 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1350 | 1374 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1382 aa Download sequence Send to blast |
MAISEQSQDV FPWLKSLPVA PEFRPTLAEF QDPIAYIFKI EEEASRYGIC KILPPVPPSS 60 KKTAINNLNR SLAARAAARV RDSGRACDYD GGPTFATRQQ QIGFCPRKQR PVQRPVWQSG 120 EHYTFGEFEF KAKSFEKNYL KKCGKKSQLS ALEIETLYWR ATVDKPFSVE YANDMPGSAF 180 IPLSLAAARR RESGGDGGTV GETAWNMRAM ARAEGSLLKF MKEEIPGVTS PMVYIAMMFS 240 WFAWHVEDHD LHSLNYLHMG AGKTWYGVPK DAAVAFEEVV RVHGYGGELN PLVTFSTLGE 300 KTTVMSPEVF VKAGIPCCRL VQNPGDFVVT FPGAYHSGFS HGEKRLFFRF NFGEASNIAT 360 PQWLRMAKDA AIRRAAINYP PMVSHLQLLY DFALALGSRV PASIHTKPRS SRLKDKKRSE 420 GEKLTKELFA QNIIHNNELL RSLGKGSPVG LLPQSSSDIS VCSDLRVGSH LSANQEKPIQ 480 LKSEDLSSDS VMVGLNNSVK DSVSVKEKFT SLCERNRNHL ASKENETHGT PTDVERRRND 540 GAVGLSDQRL FSCVTCGILS FDCVAIVQPK EAAARYLMSA DCSFFNDWTV ASGSANLGQA 600 AELLHPQSTG DHSVNYSYNV PVQTTDHSMK TGDQRTSSSS LTKAHKDNGA LGLLATAYGD 660 SSDSEEEDYK GLDIPVPEEG ACVLEASSFD TDGNEEARHG LSSAFNNERL SCAKGKEVDV 720 SHATSSCSTL SCNSEQNRLS KGGNTSLLDM TLPFIPRSDD DSSRLHVLCL EHAAEVEQQL 780 RPIGGINIML LCHPEYRRIE AEGKIVAEEL GVDHEWSDTE FRNVTREDEE TIQAALANVE 840 AKAGNSDWAV KLGINLSYSA ILSRSPLYSK QMPYNSVIYN AFGRRSPAMS SPTKPQVSGK 900 RSSRQRKYVV GKWCGKVWMS HQVHPFLLQQ DLEGEESERS CHLRVALDED VTGKRSFPYD 960 VSRDSTTMFG RKYCRKRKMR AKAVPRKKLT SFKREDGVSD DTSEDHSYKQ QWRAFGNEED 1020 MSEDSSNQMS DQQQLKEIVI HQGYKEFESD DEVSDRSLGQ EYAVRECAAS ESSMENGFQL 1080 YREKQAMSDH YDDDIDRHPR GIPRSKRTKV FRKPVSYESE ENGLYQQRGR VFTSIAQTSR 1140 MGGGYDSAEN SLEEQDFFST GKRRTRSTAK RKVKTEIVQS RRDTKGRALQ QSGSRKKNQE 1200 VGSSMEGPST RLRVRNLKPS RGSSETNPKK TDKKGRNTSF SAVASEEEVE EDEEEENEEE 1260 ECSPYQCDME GCTMSFSSEK QLSLHKRNIC PVKGCGKTFF SHKYLVQHRR VHSDDRPLKC 1320 PWKGCKMTFK WAWSRTEHIR VHTGARPYVC AEPSCGQTFR FVSDFSRHKR KTGHSVKKTK 1380 KR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 2e-76 | 1262 | 1382 | 20 | 140 | Lysine-specific demethylase REF6 |
| 6a58_A | 2e-76 | 1262 | 1382 | 20 | 140 | Lysine-specific demethylase REF6 |
| 6a59_A | 2e-76 | 1262 | 1382 | 20 | 140 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 975 | 987 | KRKMRAKAVPRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
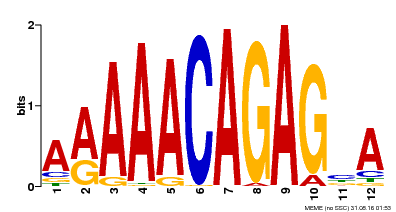
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp5g13610 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006404261.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A1J3JIY4 | 0.0 | A0A1J3JIY4_NOCCA; Lysine-specific demethylase REF6 | ||||
| STRING | XP_006404261.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5259 | 28 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



