 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp5g00890 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 333aa MW: 37127.2 Da PI: 5.1785 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.3 | 1.3e-16 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+grWT+eEd++l d+++ G g+W++ ++ g++R++k+c++rw +yl
Tp5g00890 14 KGRWTAEEDQILTDYIQSNGEGSWRALPKNAGLKRCGKSCRLRWINYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 50.8 | 3.8e-16 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+ T eE++l+v+++ lG++ W+ Ia+ ++ gRt++++k++w+++l
Tp5g00890 67 RGNITREEEDLIVKLHSSLGNR-WSMIASNLP-GRTDNEIKNYWNSHL 112
7999******************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-21 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.477 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.79E-28 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.0E-12 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.6E-14 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.19E-9 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 25.66 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-24 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.2E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-13 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.28E-11 | 71 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0051555 | Biological Process | flavonol biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 333 aa Download sequence Send to blast |
MGRAPCCEKV GMKKGRWTAE EDQILTDYIQ SNGEGSWRAL PKNAGLKRCG KSCRLRWINY 60 LRSDLKRGNI TREEEDLIVK LHSSLGNRWS MIASNLPGRT DNEIKNYWNS HLSRKLHSFI 120 RKSTVIDNAH PPQKHRPGRT RRSAMKPKFR LNPKNNKNPN SLKANESYAV LAMATKENGE 180 GGEEEDALNV LSSSSSSGAE KAGLGPCGCG NDGYCNPSIN DDDGVLCFND DIFDTCFLSN 240 EPNADHVFPS GPNKGFSSGH ENIDTMADDF VDWDFVCRDG QSLCDEKEGT DSVLSWLLDA 300 DKLESDIRQT ESNDFGEPLD LDEEKAMVAW LLS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 2e-24 | 12 | 116 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| 1mse_C | 1e-24 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 1e-24 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Modulates overall growth by reducing the proliferation activity of meristematic cells and delaying development (PubMed:18359753). Flavonol-specific transcription activator involved in the regulation of several genes of flavonoid biosynthesis. Activates the expression of CHS, CHI, F3H and FLS1 (PubMed:17419845, PubMed:20731781). Confers tolerance to UV-B (PubMed:19895401). {ECO:0000269|PubMed:17419845, ECO:0000269|PubMed:18359753, ECO:0000269|PubMed:19895401, ECO:0000269|PubMed:20731781}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
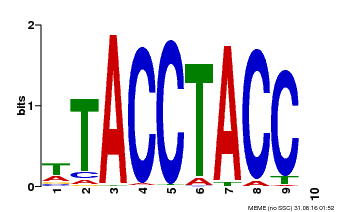
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp5g00890 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006402364.1 | 1e-173 | transcription factor MYB11 | ||||
| Swissprot | Q9LZK4 | 1e-168 | MYB11_ARATH; Transcription factor MYB11 | ||||
| TrEMBL | V4LMW4 | 1e-172 | V4LMW4_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006402364.1 | 1e-173 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G62610.1 | 1e-155 | myb domain protein 11 | ||||



