 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp3g00600 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 218aa MW: 24710.4 Da PI: 5.1754 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.1 | 3e-18 | 19 | 62 | 13 | 56 |
HHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 13 eeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+ Le+ Fe ++++ e++++LAkklgL+ rqV vWFqNrRa++k
Tp3g00600 19 HLLEKSFETENKLEPERKTQLAKKLGLQPRQVAVWFQNRRARWK 62
67*****************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 120.7 | 7.3e-39 | 18 | 100 | 11 | 93 |
HD-ZIP_I/II 11 vklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
v+lLE+sFe+e+kLeperK++la++LglqprqvavWFqnrRAR+ktkqlE+d++ Lk++yd+l ++ +++ ke+++Lr+++++
Tp3g00600 18 VHLLEKSFETENKLEPERKTQLAKKLGLQPRQVAVWFQNRRARWKTKQLERDFDLLKSTYDQLLSNYDSIVKENDKLRSQVTS 100
789****************************************************************************9975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00389 | 9.8E-16 | 6 | 68 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 6.42E-17 | 17 | 73 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 2.98E-15 | 18 | 65 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 5.2E-20 | 18 | 71 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 2.1E-15 | 19 | 62 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50071 | 15.661 | 21 | 64 | IPR001356 | Homeobox domain |
| PRINTS | PR00031 | 2.8E-6 | 35 | 44 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 39 | 62 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 2.8E-6 | 44 | 60 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 2.8E-16 | 64 | 105 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001558 | Biological Process | regulation of cell growth | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 218 aa Download sequence Send to blast |
MAKENGPHQA QKGPLQIVHL LEKSFETENK LEPERKTQLA KKLGLQPRQV AVWFQNRRAR 60 WKTKQLERDF DLLKSTYDQL LSNYDSIVKE NDKLRSQVTS LAEKLRGKDE TANEPPGQVP 120 EANQVDPVNM NRFEPAMKTE DRLSSGSVGS AVLDEDAPQL LDSCDSYFPS IVPIHSEEYN 180 NNKNNGSDND RSCFADVFVP TTSPSHDHGE PLGFWGWT |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 56 | 64 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription activator involved in leaf development. Binds to the DNA sequence 5'-CAAT[AT]ATTG-3'. {ECO:0000269|PubMed:8535134}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00323 | DAP | Transfer from AT3G01470 | Download |
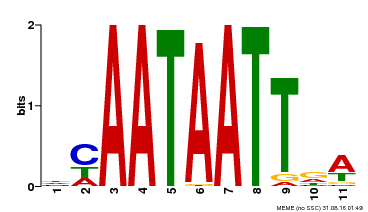
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp3g00600 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK352854 | 0.0 | AK352854.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-08-H09. | |||
| GenBank | AK353535 | 0.0 | AK353535.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-50-D05. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006408508.2 | 1e-128 | homeobox-leucine zipper protein HAT5 | ||||
| Swissprot | Q02283 | 1e-117 | HAT5_ARATH; Homeobox-leucine zipper protein HAT5 | ||||
| TrEMBL | V4NTB7 | 1e-127 | V4NTB7_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006408508.1 | 1e-128 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8884 | 27 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 1e-118 | homeobox 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



