 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp2g28610 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 491aa MW: 54669.3 Da PI: 6.5943 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 45.7 | 1.6e-14 | 227 | 286 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + g ++ ++++lg ++ +++Aa+a++ a++k++g
Tp2g28610 227 SIYRGVTRHRWTGRYEAHLWDnSCRrEGqaRKgRQVYLGGYDKEDKAARAYDLAALKYWG 286
57*******************666664478446**********99*************98 PP
| |||||||
| 2 | AP2 | 50.2 | 6.2e-16 | 329 | 380 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
Tp2g28610 329 SIYRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFATEEEAAEAYDIAAIKFRG 380
57****************988532...5************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.31E-16 | 227 | 295 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 2.0E-11 | 227 | 286 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 4.37E-22 | 227 | 296 | No hit | No description |
| PROSITE profile | PS51032 | 18.742 | 228 | 294 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 5.6E-15 | 228 | 294 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.3E-27 | 228 | 300 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.2E-6 | 229 | 240 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 2.7E-10 | 329 | 380 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 5.63E-12 | 329 | 388 | No hit | No description |
| SuperFamily | SSF54171 | 7.19E-18 | 329 | 389 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 4.0E-18 | 330 | 388 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 7.3E-31 | 330 | 394 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.177 | 330 | 388 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.2E-6 | 370 | 390 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 491 aa Download sequence Send to blast |
MEILKSPDQS HFVSYDASST PYLIDNFYVL KEEAQLEAAT MADSTTLATF FDSQTHSPVH 60 ILKLEDFLGD SSTSFARYSD NQTETQDSSS LSQIYDPRHH NVVTGFFSDH HHDLKTIACF 120 QTNSGSEIVD DSTSLGRANL TSGSEFLSVH AVESSKSAEL GIHGGCTKGG ALSLAVDNID 180 RPSSCNNGER GKNNKKTTVS KKETATEVES SDDSKKKIAE TFGQRTSIYR GVTRHRWTGR 240 YEAHLWDNSC RREGQARKGR QVYLGGYDKE DKAARAYDLA ALKYWGPTAT TNFQIASYSK 300 ELEEMNHMTK QEFIASLRRK SSGFSRGASI YRGVTRHHQQ GRWQARIGRV AGNKDLYLGT 360 FATEEEAAEA YDIAAIKFRG LNAVTNFEMN RYDVEAVMKS SFPVGGGAAK RHKLSLESPP 420 PPSDDHNLQQ DLLPSSSSDQ NPNSIPCGIP FESSVLYHHQ SFFQHYPALS DPTVQVPMNQ 480 AEFFLWPNQS Y |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00589 | DAP | Transfer from AT5G65510 | Download |
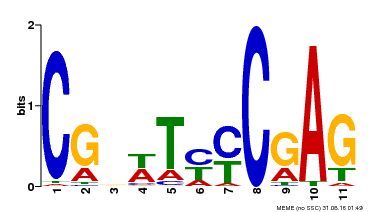
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp2g28610 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353404 | 0.0 | AK353404.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-30-G24. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024007164.1 | 0.0 | AP2-like ethylene-responsive transcription factor AIL7 | ||||
| Swissprot | Q6J9N8 | 0.0 | AIL7_ARATH; AP2-like ethylene-responsive transcription factor AIL7 | ||||
| TrEMBL | E4MY07 | 0.0 | E4MY07_EUTHA; mRNA, clone: RTFL01-30-G24 | ||||
| STRING | XP_006394021.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3333 | 27 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G65510.1 | 0.0 | AINTEGUMENTA-like 7 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



