 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp2g27280 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1063aa MW: 118753 Da PI: 6.0927 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 186.8 | 2.2e-58 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcyw 100
l+e ++rwl++ ei++iL n++k+ +++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+++vl+cyYah+e+n++fqrrcyw
Tp2g27280 19 LSEaQHRWLRPAEICEILRNHQKFLIASEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSIDVLHCYYAHGEDNENFQRRCYW 118
45559*********************************************************************************************** PP
CG-1 101 lLeeelekivlvhylevk 118
+Le++l +iv+vhylevk
Tp2g27280 119 MLEQDLMHIVFVHYLEVK 136
***************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 86.188 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.1E-86 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.2E-51 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 4.48E-14 | 467 | 553 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 20.049 | 650 | 761 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 5.13E-19 | 650 | 761 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 8.5E-8 | 651 | 726 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.2E-18 | 655 | 794 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 7.15E-16 | 656 | 759 | No hit | No description |
| SMART | SM00248 | 1500 | 667 | 696 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.351 | 667 | 699 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 11.007 | 700 | 732 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0043 | 700 | 729 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.32 | 875 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 6.9E-8 | 876 | 925 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 7.858 | 877 | 905 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0021 | 877 | 896 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0092 | 898 | 920 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.413 | 899 | 923 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.7E-4 | 901 | 920 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1063 aa Download sequence Send to blast |
MADRGSFGFA PRLDIEQLLS EAQHRWLRPA EICEILRNHQ KFLIASEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSIDVLH CYYAHGEDNE NFQRRCYWML 120 EQDLMHIVFV HYLEVKGNRM SSSGIKETNS NSLSGTASVN IDSTATPSST LSPLCEDADS 180 GDSRQASSSL QANPERQTVA PQTRHQQNAS PINSYNPTSI LGNQDGWTSA PGIGIVSQVH 240 ENRVKESDSQ RSLDVPAWDA SFDNSLARYQ NLPYNALLNQ TQTSHAGLIP MEGNKGSPLT 300 AEHLRNPLQN QVNWQIPVQD SLPLKKWPIN LHSGMSDGTD PALLGQRAHE NFGTFSSLLG 360 CQQQPIGGSF QAPFTSIEAA YIPKLGPENL LYEASANQTL PLRKSLLKKE DSLKKVDSFS 420 RWVSNELGEM EDLQMQFSSG GIAWTSVETA AAASSLSPSL SEDQRFTIID FWPKWTQTDS 480 EVEVMVVGIF LLSPQEVTNY SWSCMFGEVE VPAEILVDGV LCCHAPPHEV GQVPFYITCS 540 DRFSCSEVRE FDFLPGSTKK LNTAEIYGAF TNEASLHLRF ENLLARISSA QEHHIFEDIG 600 EKRRKISRIM LLKDEKESLL PGTIEKDLTE LETKERLIRE EFEDKLYLWL IHKVTEEGKG 660 PNILDEEGQG VLHLAAALGY DWAIKPILAA GVSINFRDAN GWSALHWAAF SGREDTVAVL 720 VSLGADSGAV TDPSPELPLG KTAADLAYGN GHRGISGFLA ESSLTSYLEK LTVDAKEHSS 780 AESSRAKAVQ TVAERTATPM GYGDVPETLS MKDSLTAVFN ATQAADRLHQ VFRMQSFQRK 840 QLSEFGDNNE FDISDELAVS FAAGKTKKPG HSGGAVHAAA VQIQKKYRGW KKRKEFLLIR 900 QRIVKIQAHV RGHQVRKQYR TIIWSVGLLE KIILRWRRKG SGLRGFKRDT VTKAPEPVCA 960 PAQEDDYDFL KEGRKQTEER LQKALTRVKS MAQYPEARAQ YRRLLTVVEG IRENEASSSS 1020 AMNNNNNNNN SNTEEAANYN EEDDLIDIDS LLDDDTFMSL AFE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
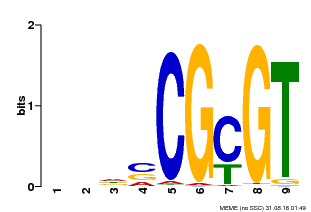
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp2g27280 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB493811 | 0.0 | AB493811.1 Arabidopsis thaliana At5g64220 mRNA for hypothetical protein, partial cds, clone: RAAt5g64220. | |||
| GenBank | AK229403 | 0.0 | AK229403.1 Arabidopsis thaliana mRNA for Calmodulin-binding transcription activator 2, complete cds, clone: RAFL16-66-C08. | |||
| GenBank | BT010874 | 0.0 | BT010874.1 Arabidopsis thaliana At5g64220 gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006394184.1 | 0.0 | calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A1J3C8U5 | 0.0 | A0A1J3C8U5_NOCCA; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A1J3JZU7 | 0.0 | A0A1J3JZU7_NOCCA; Calmodulin-binding transcription activator 2 | ||||
| STRING | XP_006394184.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



