 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp2g13490 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 603aa MW: 64619.8 Da PI: 9.3515 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.1 | 3.1e-05 | 83 | 105 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ C+k F r nL+ H r H
Tp2g13490 83 FICEVCNKGFQREQNLQLHRRGH 105
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13.1 | 0.00029 | 159 | 181 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC++C+k++ +s+ k H +t+
Tp2g13490 159 WKCEKCSKRYAVQSDWKAHSKTC 181
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 6.8E-6 | 82 | 105 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 3.71E-7 | 82 | 105 | No hit | No description |
| Pfam | PF12171 | 3.0E-5 | 83 | 105 | IPR022755 | Zinc finger, double-stranded RNA binding |
| SMART | SM00355 | 0.0052 | 83 | 105 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.99 | 83 | 105 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 85 | 105 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 110 | 124 | 154 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.8E-5 | 147 | 180 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 3.71E-7 | 154 | 179 | No hit | No description |
| SMART | SM00355 | 130 | 159 | 179 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 603 aa Download sequence Send to blast |
MAASSSSAAS FFGVRQDDQS HLLPPNSSAA APPPPPHHQP PQPQQVPLEA PPQKKKRNQP 60 RTPNSDAEVI ALSPKTLMAT NRFICEVCNK GFQREQNLQL HRRGHNLPWK LKQKSTKEVK 120 RKVYLCPEPT CVHHDPSRAL GDLTGIKKHY YRKHGEKKWK CEKCSKRYAV QSDWKAHSKT 180 CGTKEYRCDC GTLFSRRDSF ITHRAFCDAL AQESARHPTS LTSLPSHHFP YGQNTNNSNN 240 NTSSMILGLS HMGAPQNLDH QSGDVLRLGS GGGGGGGAAS RSSSDLIAAN ASGYFMQEQT 300 PGFHDQQDHH HHQHQQQGFL AASNNMKPSS MNFQQSLMQF SHDSHNSPSS NPFNLSFLSG 360 NNAVASATSN PNAGAVSSGN LMISNHFDGE NAVGGGGGGE GSTGLFPNNL MSSADRISSG 420 AVPSLFSSSM QNPNSTPHMS ATALLQKAAQ MGSTSSNNNN NGSNNSNNNN ASSILRSFGS 480 GMYGENESNL HDLMNSFSNP GATGNNVNGV DSPFGSYGGV NKGLNADKQS MTRDFLGVGQ 540 IVRSMSGSGG FQQQQQQQQQ QHGNNSRERV GSSSDSADRN SMNVNPGGGP ASSPPYGIHH 600 ASF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 8e-34 | 155 | 216 | 3 | 64 | Zinc finger protein JACKDAW |
| 5b3h_F | 8e-34 | 155 | 216 | 3 | 64 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a positive regulator of the starch synthase SS4. Controls chloroplast development and starch granule formation (PubMed:22898356). Binds DNA via its zinc fingers (PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). {ECO:0000269|PubMed:22898356, ECO:0000269|PubMed:24821766}. | |||||
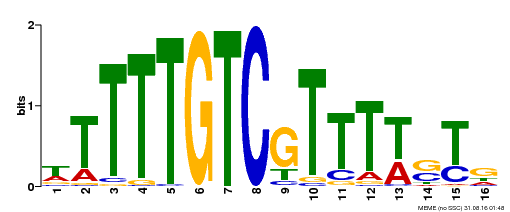
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00255 | DAP | Transfer from AT2G02070 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp2g13490 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HM222924 | 0.0 | HM222924.1 Thellungiella parvula phosphatase, cyclin J18, clathrin assembly protein-related protein, BPC1, SGR5/ATIDD15, leucine-rich receptor kinase, endomemebrane protein 70, salt overly sensitive 1, glutamate decarboxylae 4, oligopeptide transport protein, peptide transporter, and NADH-ubiquinone oxidoreductase B18 genes, complete cds; and unknown genes. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009123129.1 | 0.0 | PREDICTED: protein indeterminate-domain 5, chloroplastic-like | ||||
| Swissprot | Q9ZUL3 | 0.0 | IDD5_ARATH; Protein indeterminate-domain 5, chloroplastic | ||||
| TrEMBL | A0A397Y3X1 | 0.0 | A0A397Y3X1_BRACM; Uncharacterized protein | ||||
| STRING | Bra040974.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4373 | 26 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G02070.1 | 0.0 | indeterminate(ID)-domain 5 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



