 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp1g32340 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 205aa MW: 23216.8 Da PI: 7.0239 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 61.8 | 1.5e-19 | 80 | 129 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkr..krfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+W+AeIrdp + r++lg+f tae Aaka++aa+++++g
Tp1g32340 80 RYRGVRQRP-WGKWAAEIRDP-----HkaMRVWLGTFETAEAAAKAYDAAALRFRG 129
7********.**********7.....337************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.96E-22 | 80 | 139 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 4.31E-22 | 80 | 139 | No hit | No description |
| PROSITE profile | PS51032 | 25.58 | 80 | 137 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 7.6E-33 | 80 | 138 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.5E-37 | 80 | 143 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 4.4E-13 | 80 | 129 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.3E-11 | 81 | 92 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.3E-11 | 103 | 119 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0034605 | Biological Process | cellular response to heat | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 205 aa Download sequence Send to blast |
MVSMLTHVVS GETEPSASVR RFGASSSSGS MSHCHKREQE EFHLHRPLVT TELVDTSSIT 60 GSSVAQEYES STKSLEKPRR YRGVRQRPWG KWAAEIRDPH KAMRVWLGTF ETAEAAAKAY 120 DAAALRFRGS KAKLNFPENV GTQTIQQDSH YLQNPLQPPT TYLDQCPSLE QQPPLVGMLQ 180 STEEERHFFT QPWTDYDPYH YLSSG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 2e-24 | 77 | 142 | 2 | 68 | ATERF1 |
| 3gcc_A | 2e-24 | 77 | 142 | 2 | 68 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in the regulation of plant development and tolerance to abiotic stresses (PubMed:21069430). Binds to the GCC-box pathogenesis-related promoter element and the cis-element CE1 (coupling element 1). Involved in the regulation of gene expression in response to abiotic stresses, possibly through the abscisic acid (ABA) signaling pathway (PubMed:20193749). Involved in resistance to the beet cyst nematode Heterodera schachtii in roots. May promote callose deposition at syncytia which may interfere with nutrient import into syncytia and inhibit the development of nematodes (PubMed:23510309). {ECO:0000269|PubMed:20193749, ECO:0000269|PubMed:21069430, ECO:0000269|PubMed:23510309}. | |||||
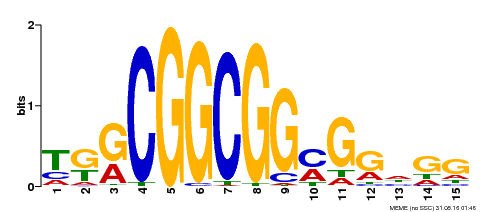
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00183 | DAP | Transfer from AT1G43160 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp1g32340 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salt, heat and drought stresses (PubMed:21069430). Induced by osmotic stress (PubMed:20193749). Induced by jasmonate (JA) (PubMed:21069430, PubMed:14756769, PubMed:17786451). Induced by salicylic acid (SA) (PubMed:14756769, PubMed:21069430). Induced by abscisic acid (ABA) (PubMed:20193749, PubMed:21069430). Induced by ethylene (PubMed:14756769). Induced by infection with the bacterial pathogen Pseudomonas syringae pv. tomato DC3000 (PubMed:14756769, PubMed:23510309). Induced by the bacterial pathogen Pseudomonas syringae pv. maculicola ES4326 (PubMed:14756769). Induced by wounding (PubMed:17786451). Down-regulated by infection with the beet cyst nematode Heterodera schachtii (PubMed:23510309). {ECO:0000269|PubMed:14756769, ECO:0000269|PubMed:17786451, ECO:0000269|PubMed:20193749, ECO:0000269|PubMed:21069430, ECO:0000269|PubMed:23510309}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_175008.1 | 1e-82 | related to AP2 6 | ||||
| Swissprot | Q7G1L2 | 1e-83 | RAP26_ARATH; Ethylene-responsive transcription factor RAP2-6 | ||||
| TrEMBL | A0A178W859 | 4e-82 | A0A178W859_ARATH; RAP2.6 | ||||
| STRING | AT1G43160.1 | 5e-82 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10 | 28 | 1650 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G43160.1 | 2e-81 | related to AP2 6 | ||||



