 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp1g26030 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 334aa MW: 37049.4 Da PI: 8.108 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 109.9 | 3.6e-34 | 46 | 162 | 2 | 135 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssaseceaesssssasnsssgkaaksaakskksq 101
g+kdrhsk+ T++g+RdRR+Rls+ +a++f+dLqd+LGfd++sk++eWL+++a +i +l+ +++ + + s +++ +s
Tp1g26030 46 SGGKDRHSKVLTSKGLRDRRIRLSVATAIQFYDLQDRLGFDQPSKAVEWLINAASDSISDLPLINTNFDQ-----TLS-------------LSKSACSSG 127
689**********************************************************999555541.....122.............222233455 PP
TCP 102 ksaasalnlak.esrakarararertrekmriknk 135
+s++s l+l++ e r kar+rarert++++ ++ +
Tp1g26030 128 TSESSLLSLSRtEIRGKARERARERTAKEKDKDLQ 162
6666679999999***********99998877654 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 9.2E-33 | 47 | 160 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 31.035 | 48 | 106 | IPR017887 | Transcription factor TCP subgroup |
| PROSITE profile | PS51370 | 9.766 | 139 | 157 | IPR017888 | CYC/TB1, R domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MEVDDDIDAQ QQTSRKLQRF SSDDNNTGMR DWSNPSSRII RVSRASGGKD RHSKVLTSKG 60 LRDRRIRLSV ATAIQFYDLQ DRLGFDQPSK AVEWLINAAS DSISDLPLIN TNFDQTLSLS 120 KSACSSGTSE SSLLSLSRTE IRGKARERAR ERTAKEKDKD LQNAQSSSFT QLLTGGFDEP 180 NRHWTGGSSD CFNPVQLIQN SSSSSLPQFN NHRQESSLNH HNQSQFSFVP DYNFGISSSD 240 SPGATTINGG CYSSRGTLQS NSQSLLLNNI NQRSISSSSS SSSPMDSQSI SFFMATPPPL 300 DHHSHQLPAT FDGRLYLYYG EGNRNTEDKG KDRR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 1e-20 | 53 | 107 | 1 | 55 | Putative transcription factor PCF6 |
| 5zkt_B | 1e-20 | 53 | 107 | 1 | 55 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Plays a pivotal role in the control of morphogenesis of shoot organs by negatively regulating the expression of boundary-specific genes such as CUC genes, probably through the induction of miRNA (e.g. miR164). In association with ABAP1, exerts a negative role in cell proliferation in leaves, possibly by inhibiting mitotic DNA replication. Participates in ovule develpment (PubMed:25378179). {ECO:0000269|PubMed:17307931, ECO:0000269|PubMed:18818695, ECO:0000269|PubMed:25378179}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00169 | DAP | Transfer from AT1G30210 | Download |
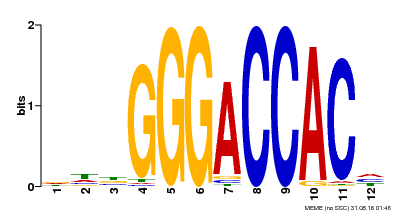
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp1g26030 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353156 | 0.0 | AK353156.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-21-I20. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024008427.1 | 1e-180 | transcription factor TCP24 | ||||
| Refseq | XP_024008428.1 | 1e-180 | transcription factor TCP24 | ||||
| Swissprot | Q9C758 | 1e-151 | TCP24_ARATH; Transcription factor TCP24 | ||||
| TrEMBL | E4MXA9 | 1e-178 | E4MXA9_EUTHA; mRNA, clone: RTFL01-21-I20 | ||||
| STRING | Bra032365.1-P | 1e-179 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM13840 | 17 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G30210.2 | 1e-121 | TCP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



