 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010548015.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 419aa MW: 45646.3 Da PI: 9.5183 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 44.2 | 4.2e-14 | 341 | 384 | 5 | 48 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkke 48
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++ N +L+k+
XP_010548015.1 341 RRQRRMIKNRESAARSRARKQAYTMELEAEVAKLKELNNELQKK 384
69*************************************99986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 2.2E-12 | 337 | 402 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.542 | 339 | 384 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 3.16E-10 | 341 | 389 | No hit | No description |
| Pfam | PF00170 | 5.1E-12 | 341 | 385 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 4.70E-24 | 341 | 395 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 7.0E-14 | 341 | 388 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 344 | 359 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 419 aa Download sequence Send to blast |
MGSRVNFRNF ADGDMNQKQA VIGNVPLARQ NSVFSLTFDE FQNTWVGLGK DFGSMNMDEL 60 LKNIWTAEES QAMNTTTTTT NNINNGNNSN IGNSNGLAVA MGGGGDGCVG FSGGNLQRQG 120 SFTLPRTLSQ KTVDEVWREL MKDSDNGGGG CSNGGSDVPQ RQQTLGEITL EEFLVRAGVV 180 REEPQPLERP DNINGGFYGF GGNASLGLHQ NNFVSNHTLD FSGNAAGVRL GGLPPRSQPQ 240 QETAFPKQAT IAFTNNVDMA NISKPIIQEV RSTIAGVQDC PVKSNLLQAV NLKTGVTVAA 300 VSPGGQMSPD LTPRDASVSP VPYMFGRGRR SGAALEKVIE RRQRRMIKNR ESAARSRARK 360 QAYTMELEAE VAKLKELNNE LQKKQAVIME VQKNQLLEPI RQPWGCKRQC LRRTLTGPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
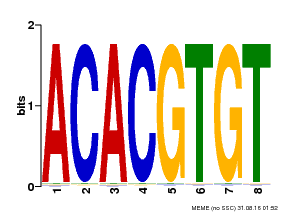
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010548015.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010548013.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Refseq | XP_010548014.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Refseq | XP_010548015.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Refseq | XP_010548017.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Swissprot | Q9M7Q3 | 0.0 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | E4MY11 | 0.0 | E4MY11_EUTHA; mRNA, clone: RTFL01-30-J01 | ||||
| TrEMBL | V4MBG2 | 0.0 | V4MBG2_EUTSA; Uncharacterized protein | ||||
| STRING | XP_010548013.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34000.2 | 1e-149 | abscisic acid responsive elements-binding factor 3 | ||||



