 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010547792.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 860aa MW: 95683.4 Da PI: 6.8391 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74.8 | 9.8e-24 | 166 | 267 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrse 89
f+k+lt sd++++g +++ +++a+e+ + + + +++l+ +d++g++W++++i+r++++r++l++GW+ Fv++++L +gD ++F + ++
XP_010547792.1 166 FCKTLTASDTSTHGGFSVLRRHADEClppldmSRQ-PPTQELVAKDLHGNEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGEN 257
99*****************************7333.34459************************************************..4499 PP
E..EEEEE-S CS
B3 90 felvvkvfrk 99
+el+v+v+r+
XP_010547792.1 258 GELRVGVRRA 267
99*****996 PP
| |||||||
| 2 | Auxin_resp | 117.3 | 1.3e-38 | 292 | 374 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha tk++F v+Y+Pr+s+seF+v+++++++++k+++s+GmRfkm+fe+e+++e+r++Gtvvg++d+dp rW nSkWr+Lk
XP_010547792.1 292 AWHAYLTKTMFIVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTVVGIEDSDPKRWANSKWRCLK 374
799******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 2.9E-41 | 162 | 281 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.4E-40 | 162 | 295 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 7.90E-22 | 164 | 266 | No hit | No description |
| PROSITE profile | PS50863 | 12.012 | 166 | 268 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.0E-23 | 166 | 268 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.2E-21 | 166 | 267 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 2.0E-35 | 292 | 374 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 7.2E-11 | 720 | 819 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 4.46E-13 | 732 | 811 | No hit | No description |
| PROSITE profile | PS51745 | 22.773 | 735 | 818 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 860 aa Download sequence Send to blast |
MATSEISTKG NSIHVRGEGF SSGYSDPTEA RNAASAGGGG GEGQQSHSTR SATVDPEAAL 60 FKELWHACAG PLVTVPKKDD RVFYFPQGHI EQVEASMSQR TEQQMPVYEL PSKILCRVIN 120 VELKAEPDTD EVFAQITLLP ESNQDENAVE KLPPPPPPPR FQVHSFCKTL TASDTSTHGG 180 FSVLRRHADE CLPPLDMSRQ PPTQELVAKD LHGNEWRFRH IFRGQPRRHL LQSGWSVFVS 240 SKRLVAGDAF IFLRGENGEL RVGVRRAMKQ QGNVPSSVIS SHSMHLGVLA TAWHAYLTKT 300 MFIVYYKPRT SPSEFIVPFD QYMESVKNNY SIGMRFKMRF EGEEAPEQRF TGTVVGIEDS 360 DPKRWANSKW RCLKVRWDET SNIPRPDRVS PWKIEPALAP PALSPVPMPR PKRPRANIVP 420 SSPDSSVLTR DGSSKATMDP LPASGLSRVL QGQEYSTLRG KITDSAECGA PENSSIWPPS 480 VAEEKVDMVC TSRRYESENW MPSGRHEPTY TDLLSGFGSN VDTSCSHRLP FTDHSAPAPA 540 PTKKMLGDQD GKFNYLGNHW PLMHSGLSLK LPESSKVPVV GDASFQGHGN TKYGGCSEYP 600 VLQFQTVENA RGNWPMPSRG LNYFDDMVQA RELAAKQPVT IQEKETGKSK DGNCRLFGIP 660 LVSSHLNETE PTVSQRNKMD EQVPQIQNLS PKVQDQSDRS KGSKSMNDHR AEQGRQPQTS 720 HAHSKDAHGK SNSTRSCTKV HKQGIALGRS VDLSKFHDYD ELVAELDRLF EFNGELMATK 780 DWLIVYTDDE GDMMLVGDDP WQEFCCMVRK IFIYTKEEVR KMNPGALNAR SEEDTAVAEG 840 YDAKDAKSVS NPSVSSTENC |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-174 | 59 | 396 | 18 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-174 | 59 | 396 | 18 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-174 | 59 | 396 | 18 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-174 | 59 | 396 | 18 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-174 | 59 | 396 | 18 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
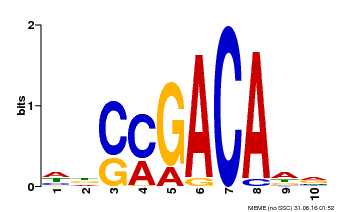
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010547792.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010547792.1 | 0.0 | PREDICTED: auxin response factor 2-like | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A1J3I677 | 0.0 | A0A1J3I677_NOCCA; Auxin response factor | ||||
| STRING | XP_010547792.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5348 | 26 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||



