 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010542800.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 748aa MW: 81904.6 Da PI: 6.6981 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.9 | 3.2e-15 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT++E+ ++++a +++G W +I +++g ++t+ q++s+ qk+
XP_010542800.1 24 RERWTEDEHSRFLEALRLYGRA-WQRIEEHIG-TKTAVQIRSHAQKF 68
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.39E-15 | 18 | 72 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.323 | 19 | 73 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-7 | 21 | 70 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.1E-17 | 22 | 72 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.0E-11 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.8E-12 | 24 | 67 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.54E-8 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 748 aa Download sequence Send to blast |
MEKNISGEEL VTKIRKPYTI TKQRERWTED EHSRFLEALR LYGRAWQRIE EHIGTKTAVQ 60 IRSHAQKFFT KQPYAWKAST WMRFAVDASI ELHKKLLEKE AQAKGIPICQ ALDIEIPPPR 120 PKRKPSNPYP RKTGNGESTS QVSSAKVVKL DSSASFSHCN QALDLEKKPF TEKPNGVEKV 180 PNGNGNQDDN CSGVSARPTE AHCSSISSVN KYPFPNKAAP RNPSSFRESE PDLTKATADN 240 GTSKTSNADN SCYSQDNDAN VLPHISISKE DNGVNGLSPV GNMNGIQSYP WHFPVHVLNG 300 NVGTYPQNPP GMGFQDFMFR PVGEVHGHSG LFTIPSTSAT SQHQDNVPRS SRQAFPAFHP 360 AFVPACHTQD DYRSFLQISS TFSNLIMSAL LQNPAAHAAA TFAASFWPYG NVGSSNDSST 420 STTIQTDGAF PPRQMSSPPP SMAAITAATV AAATAWWAAH GLLPICAPFT CLPMSAPIVP 480 PSVSAQTELP ETDKKDAATA NPQQPAMEQG EAQVLASPFE SDDDLDENGR TEQDTGLKTE 540 DHKEKVVEVQ EQVQDSNTTQ KKKQVDRSSC GSNTPSGSDV ETDALEKAAE KDKEEDVKET 600 DGNQTGTESS TRRTSRSKDN SSDSWKEVSE EGRMAFDALF ARERLPQSFS LALDVEKESP 660 EKGKGNREER EREMAHKSKI QDSCSKETAS RSEGNGGITI GIGEGKSLRA RRTGFKPYKR 720 CSMEAKESSS QAEEKVSKRI RLDSKAST |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the circadian clock. Binds to the promoter region of APRR1/TOC1 and TCP21/CHE to repress their transcription. Represses both CCA1 and itself. {ECO:0000269|PubMed:12015970, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:9657154}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
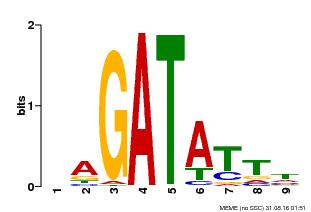
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010542800.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation with peak levels occurring around 1 hour after dawn. Up-regulated by APRR1/TOC1 and transiently by light treatment. Down-regulated by APRR5, APRR7 and APRR9. {ECO:0000269|PubMed:12574129, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:19286557, ECO:0000269|PubMed:20233950, ECO:0000269|PubMed:9657154}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010542800.1 | 0.0 | PREDICTED: protein LHY-like isoform X1 | ||||
| Refseq | XP_010542801.1 | 0.0 | PREDICTED: protein LHY-like isoform X1 | ||||
| Refseq | XP_010542802.1 | 0.0 | PREDICTED: protein LHY-like isoform X1 | ||||
| Swissprot | Q6R0H1 | 0.0 | LHY_ARATH; Protein LHY | ||||
| TrEMBL | A0A061DT31 | 0.0 | A0A061DT31_THECC; Homeodomain-like superfamily protein isoform 1 | ||||
| STRING | XP_010542800.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5576 | 26 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01060.4 | 1e-144 | MYB_related family protein | ||||



