 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010541362.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 479aa MW: 52846.7 Da PI: 7.4116 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 55.6 | 1.3e-17 | 161 | 210 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV++++++grW+++I+d + k+++lg f+ta Aa+a+++a+ k++g
XP_010541362.1 161 SQYRGVTFYRRTGRWESHIWD------CgKQVYLGGFDTAHAAARAYDQAAIKFRG 210
78*******************......55************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.60E-13 | 161 | 220 | No hit | No description |
| SuperFamily | SSF54171 | 6.54E-16 | 161 | 220 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 9.5E-10 | 161 | 210 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 9.1E-32 | 162 | 224 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.2E-17 | 162 | 218 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 17.899 | 162 | 218 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 479 aa Download sequence Send to blast |
MLDLNLEVGS AESIQEGADS ARDKGISGVV SNQMDESVTS NSSVVNADGT NCVDGEEESC 60 STRAVTFHFD ILKGGDREGE AEKEVVMTKE LFPVTGGNGN GRGCLISEQQ SCKNSVDISF 120 QSWKQAGEIL GHGVGGGAKR EMQQTSRPVK KSRRGPRSKS SQYRGVTFYR RTGRWESHIW 180 DCGKQVYLGG FDTAHAAARA YDQAAIKFRG LDADINFSVS DYEDDTKRMA NLTKEEFVQI 240 LRRQSTGFSR GGSRYQGAKF GRAYDKAAIE WNERNATANI EARAYQMIPE ASNRAHVKLD 300 LNLGISPPVG NYPQQNERSL QLHRDHTDIV GGWDSKVHSP TYTSLLYFIL ENRMAVAVHN 360 PPFDFLKRGS DHPIPWNASP YPCFSAEERK LEEGIVESRS RQQTFPTRIW QAQSQPLLGN 420 AASSGFSVSA TPFSSTASFL PPRPFSNLTL FGLYLASPST AGNTSHPYHH LMQPPHPPP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). Regulates negatively the transition to flowering time and confers flowering time delay. {ECO:0000250, ECO:0000269|PubMed:14555699}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00616 | PBM | Transfer from AT5G60120 | Download |
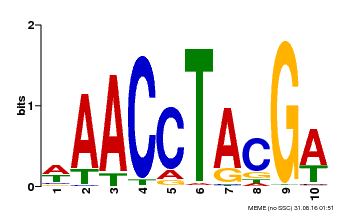
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010541362.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by miR172a-2/EAT. {ECO:0000269|PubMed:14555699}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010541361.1 | 0.0 | PREDICTED: AP2-like ethylene-responsive transcription factor TOE2 isoform X1 | ||||
| Refseq | XP_010541362.1 | 0.0 | PREDICTED: AP2-like ethylene-responsive transcription factor TOE2 isoform X1 | ||||
| Swissprot | Q9LVG2 | 1e-148 | TOE2_ARATH; AP2-like ethylene-responsive transcription factor TOE2 | ||||
| TrEMBL | E4MWP9 | 1e-156 | E4MWP9_EUTHA; mRNA, clone: RTFL01-19-B14 | ||||
| TrEMBL | V4LIS4 | 1e-156 | V4LIS4_EUTSA; Uncharacterized protein | ||||
| STRING | XP_010541361.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G60120.2 | 1e-138 | target of early activation tagged (EAT) 2 | ||||



