 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010534614.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 409aa MW: 45646.7 Da PI: 8.6151 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102 | 3.7e-32 | 80 | 133 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtpeLH++F++ave+LGG+++AtPk +l++m+vkgL+++hvkSHLQ+YR+
XP_010534614.1 80 PRLRWTPELHRCFLQAVERLGGQDRATPKLVLQTMNVKGLSIAHVKSHLQMYRS 133
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-30 | 76 | 134 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.199 | 76 | 136 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.97E-16 | 77 | 135 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.3E-24 | 80 | 135 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.2E-9 | 81 | 132 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 409 aa Download sequence Send to blast |
MRSSTNSSEN SKTCPSNSCK TIKKEEEDKD IDDEGDQEEA KTRGDHQSSS SNSYVEECPG 60 HHQIKKNGGS VRPYNRSKTP RLRWTPELHR CFLQAVERLG GQDRATPKLV LQTMNVKGLS 120 IAHVKSHLQM YRSKKADEPN QGQQGFSFET GTGYTCSLSQ VPMLQSFDQR PSTSLGYDDP 180 SWSDHGPLIN CNPWRGLTTR DGARSGQPLF ISLNERFHRS SNMGTFGIGS HSSSKKTTFN 240 LNNEEVEAFQ RSHRRTLEDL KIFNGSWVRS LCDPSSTEKW IDVSTHRGKI ATSNSVRVAE 300 GSGIVLKRKR LFLFEDCDHN KSCQDLDLSL SLKATPQTYA NVGLKGKGCL LFGDEHGQDH 360 QDCKSLSLSL SSSSSSRLSL STPTKEDDLR DHNRKRKISV SASTLDLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 3e-18 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 3e-18 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 3e-18 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 3e-18 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
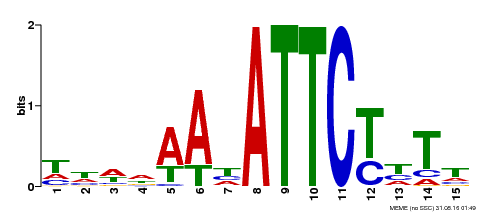
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010534614.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010534614.1 | 0.0 | PREDICTED: uncharacterized protein LOC104810118 | ||||
| TrEMBL | D7LEH7 | 1e-118 | D7LEH7_ARALL; Uncharacterized protein | ||||
| STRING | XP_010534614.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9359 | 21 | 36 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 1e-118 | G2-like family protein | ||||



