 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010532504.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 844aa MW: 97072.4 Da PI: 7.3791 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 83.4 | 3.7e-26 | 88 | 190 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerr....traetrtgCkaklkvkkekdgkwevt 79
+fY+eYA+ +GF++ +++s++sk+++e+++++f Cs++g+++e +k+ + +ra +t+Cka+++vk++ dgkw ++
XP_010532504.1 88 SFYQEYARAMGFNTAIQNSRRSKTTREFIDAKFACSRYGTKREYDKSfnkprvsqnkqeP----EnmagRRACGKTDCKASMHVKRRPDGKWIIH 178
5*******************************************99999**988754433....2345589999********************* PP
FAR1 80 kleleHnHelap 91
+++ eHnHel p
XP_010532504.1 179 SFVREHNHELLP 190
*********975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 5.6E-24 | 88 | 190 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 3.4E-26 | 288 | 380 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.446 | 568 | 604 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 2.5E-4 | 577 | 603 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.7E-8 | 579 | 606 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 844 aa Download sequence Send to blast |
MDINLRLPSG DQSKEDGEER GIDNVLEDEH KLHNGDMEIG KMEDVTVEVQ ADDSVGMVPT 60 AELVEYKEGM NLEPLNGMEF ESHSEAYSFY QEYARAMGFN TAIQNSRRSK TTREFIDAKF 120 ACSRYGTKRE YDKSFNKPRV SQNKQEPENM AGRRACGKTD CKASMHVKRR PDGKWIIHSF 180 VREHNHELLP AQAVSEQTRK IYAAMARQFA EYKTVISLNN NTKSSFEKGR NSAVETGDIK 240 ILLDFLSRMQ SMNSNFFYAV DLGDDQQLKT VFWVDAKSRH DYGSFCDVVS LDTTYVRNKY 300 RMPLAIFVGV NQHYQFMVLG CAVISDESAT TFSWIMHTWL KAMGGLAPKV IITEQDSVMN 360 SVVPEVFPNT RHCFFLWHVL VKVSENLGQV IKQHDNFTPK FEKCIYKSWR DEDFARRWWK 420 ILDRFGLKED QWMQSLYEDC RKWVPAYMTD VLLAGMSTSQ RTDSINAFFD KYLHKKTSVQ 480 EFVKLYDAIL QDRYEEEAKA DSDAWNKQPT MKSPSPFEKS VSEIYTPAVF RKFQVEVLGA 540 IACSPREENR DGTCFIFRVQ DFEKNQDFVV TWNQTKSQVS CICRLFEYKG YLCRHALNVL 600 QRCHLSSIPS RYILKRWTKD AKNRDFTGEP ERLQTRLQRY NDLCRRALKL SEEASFSQES 660 YNIAYRAIEE ALENCAGVNT AGKSPQELVT SPTQGLISVE EDNQSRSTGK TSKKKNPTKK 720 RKVNSEQEVM PVGAPENMQQ MDKLSPRTVS IESYYATQPS VQGMVQLNLM APTRESFYGN 780 QQIQGLGQLN SIASSHGSYY SAQQGIPEMS GLDFFRPSAF SYDIRDDPNV RTAQLHNNAS 840 RHPP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
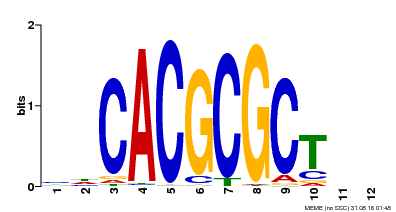
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010532504.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010532502.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_010532503.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_010532504.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_019057594.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_019057595.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A1J3CXP8 | 0.0 | A0A1J3CXP8_NOCCA; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| STRING | XP_010532502.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



