 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010529652.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 934aa MW: 102930 Da PI: 5.174 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 91.9 | 4.8e-29 | 587 | 637 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isl++l++yF +++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rkik++
XP_010529652.1 587 EKTISLDVLQQYFTGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKIKKV 637
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF55781 | 8.34E-5 | 210 | 281 | IPR029016 | GAF domain-like |
| PROSITE profile | PS51519 | 17.746 | 576 | 657 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 7.1E-26 | 589 | 637 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 1.96E-22 | 835 | 922 | No hit | No description |
| PROSITE profile | PS51745 | 23.4 | 838 | 920 | IPR000270 | PB1 domain |
| SMART | SM00666 | 4.8E-25 | 838 | 920 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 2.2E-25 | 838 | 919 | No hit | No description |
| Pfam | PF00564 | 2.7E-18 | 839 | 919 | IPR000270 | PB1 domain |
| CDD | cd06407 | 6.43E-35 | 839 | 919 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 934 aa Download sequence Send to blast |
MCEPDEDSTR NGAIPPLPRP RELLMDLDDL DLDGSWPLDQ IPHLFSSNQF ISPFVVSSSS 60 EQPFSPLWAF SDGGGTAAVP TAGDDEKLSP NSGAPPFRLA EYPIFLPCAA SSKVENTAER 120 DRNVQFPSPL TGLAPPENTD NYCVIKERMT QALRYFKEST EQQILAQVWA PVRKNGRELL 180 TTLGQPFVLN PNGNGLNQYR MISLTYMFSV DSESDIELGL PGRVYRQKLP EWTPNVQYYS 240 SREFSRLDHA LHYNVRGTLA LPVFNPSGQS CIGVVELIMT SEKIHYAPEV DRVCKALEAV 300 NLKSSEILDH QTTQICNEGR QNALAEILEI LTVVCETYKL PLAQTWVPCQ HRSVLANGGG 360 LKKSCTSFDG SCMGQVCMST TDMAAYVVDA HVWGFRDACL EHHLQKGQGV AGRAFLNRSS 420 CFCRDVTKFC KIQYPLVHYA LMFKLTSCYA ICLQSSHTGD DDYILEFFLP PAITDKSEQD 480 SLLGSLLATM KQHFQSLRVA SGISFGEEAN EVSVEFIEAL PDKKLNSRIE SIHVPSSGFE 540 SNVGEMLLDP QPVVPSSEPV KETSNVADDN GVVKEMKKPE RKRGKTEKTI SLDVLQQYFT 600 GSLKDAAKSL GVCPTTMKRI CRQHGISRWP SRKIKKVNRS ITKLKRVIES VQGTDGALDL 660 TSMAVGSIPW TNGQATAQPL NSPSGYKEPE PPKNNDSTHN GSNGHSPHEP HGSPPSNGCK 720 PSRTRDDSVG TPTSHGSCEG SYRDEMKALN PEPIFMADVS AASFSMPNRI LGSIEPFRGM 780 PIEESGSSKD LRNLCHSVAA DEKFPESNWL NNGNNIYVRP EEEAMEITRV ASGSETRSVT 840 IKASYKEDII RFRISSSSGL MELKEEIAKR LKLDAGTFDI KYLDDDSEWV LIACDVDLQE 900 CLDISGFSGS SLVRLLVQDV MANPGSSCES TGEL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
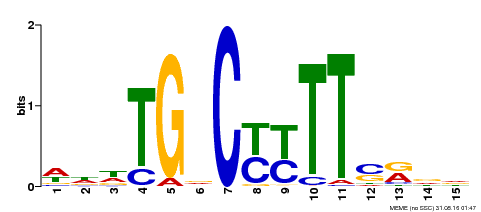
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010529652.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010529652.1 | 0.0 | PREDICTED: protein NLP7-like isoform X1 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | V4MHY8 | 0.0 | V4MHY8_EUTSA; Uncharacterized protein | ||||
| STRING | XP_010529652.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



