 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG044611t3 | ||||||||
| Common Name | TCM_044611 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 541aa MW: 58721.4 Da PI: 4.6855 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.6 | 2.5e-17 | 39 | 86 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +l+++vk++G g+W+++ ++ g+ R++k+c++rw ++l
Thecc1EG044611t3 39 KGPWTSAEDAILIEYVKKHGEGNWNAVQKHSGLFRCGKSCRLRWANHL 86
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 54 | 3.9e-17 | 92 | 135 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE++l++++++++G++ W++ a++++ gRt++++k++w++
Thecc1EG044611t3 92 KGAFTQEEEQLIIELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 135
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.409 | 34 | 86 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.8E-30 | 37 | 133 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.2E-15 | 38 | 88 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.2E-24 | 40 | 93 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.92E-12 | 41 | 86 | No hit | No description |
| Pfam | PF13921 | 4.1E-16 | 42 | 102 | No hit | No description |
| PROSITE profile | PS51294 | 26.872 | 87 | 141 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.3E-17 | 91 | 139 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-26 | 94 | 140 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.05E-12 | 94 | 135 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 541 aa Download sequence Send to blast |
MSHTKNERED GMLSEDQAES PLADDGNCGG AGGGVVMKKG PWTSAEDAIL IEYVKKHGEG 60 NWNAVQKHSG LFRCGKSCRL RWANHLRPNL KKGAFTQEEE QLIIELHAKM GNKWARMAAH 120 LPGRTDNEIK NYWNTRIKRR QRAGLPLYPP EVCLQALQES HSTSAVNGGD KGPQDIMQNN 180 SYQIPDVIFD SLKANQSVLP YVPELPDIST SSMLMKGLGS SQYCSFMPPT IHRQKRLRES 240 TAFFPGYTGA VKNECPLFDQ FQDDMSDKAA QSFGVSFQIE PDPTAKNSES FGVFPGSHAL 300 TNGNFSASES SLEAVKLELP SLQYPETELG NWGTISCPPP LLESVDAFIQ SPPPTSGVES 360 DSLSPHNSGL LDALLHEAKT LSSAKNHASD KSSNSSTPGD IAESSNFNIC ETEWENCGQP 420 LSPMGHSATS LFSECISASG SSLDEQPPAE TFTEIQKEAP IRLDSTRPDT LLASNWLEQG 480 CVYDKDQTIM SDAIATLLGD DLSSEYKNMA AGTSISSQAW GLGSCAWNNM PAVCQMSELP 540 * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-32 | 39 | 140 | 7 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of alpha-amylase expression that binds to 5'-CAACTGTC-3' motif in target gene promoter (PubMed:11743113). Positive regulator of abscisic acid (ABA) responses leading to growth arrest during seed germination (PubMed:17217461). In vegetative tissues, inhibits growth by reducing cell proliferation. Promotes the expression of aleurone-related genes (e.g. CP1, CP, GASA1, BXL1 and BXL2) in seeds. Together with MYB65 and MYB101, promotes the programmed cell death (PCD) the vacuolation of protein storage vacuoles (PSVs) in the aleurone layers during seed germination (PubMed:20699403). Binds to a GARE site (GA-response element) in the LEAFY promoter, essential for its gibberellic acid (GA)-mediated induction (PubMed:15226253). Together with MYB65, facilitates anther and tapetum development (PubMed:15722475). {ECO:0000269|PubMed:11743113, ECO:0000269|PubMed:15226253, ECO:0000269|PubMed:15722475, ECO:0000269|PubMed:17217461, ECO:0000269|PubMed:20699403}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
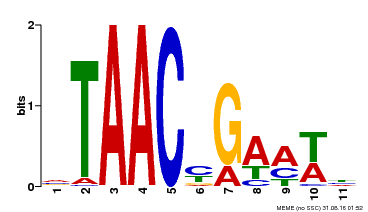
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Accumulates at the shoot apex upon the transition from short- to long-day photoperiods leading to flowering and after gibberellins (GAs) treatment (PubMed:11743113). Repressed by microRNA159 (miR159a and miR159b) in vegetative tissues (PubMed:15226253, PubMed:20699403, PubMed:17916625). Specific expression in floral organs and in the shoot apices is regulated via miR159-mediated degradation (PubMed:15722475). Repressed in germinating seeds by miR159-mediated cleavage in an abscisic acid (ABA) and ABI3-dependent manner, probably to desensitize hormone signaling during seedling stress responses (PubMed:17217461, PubMed:18305205). Slightly induced by ethylene and cytokinins (PubMed:9839469). {ECO:0000269|PubMed:11743113, ECO:0000269|PubMed:15226253, ECO:0000269|PubMed:15722475, ECO:0000269|PubMed:17217461, ECO:0000269|PubMed:17916625, ECO:0000269|PubMed:18305205, ECO:0000269|PubMed:20699403, ECO:0000269|PubMed:9839469}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007010686.2 | 0.0 | PREDICTED: transcription factor GAMYB | ||||
| Refseq | XP_007010687.2 | 0.0 | PREDICTED: transcription factor GAMYB | ||||
| Swissprot | Q8W1W6 | 1e-152 | MYB33_ARATH; Transcription factor MYB33 | ||||
| TrEMBL | A0A061FRF4 | 0.0 | A0A061FRF4_THECC; Myb domain protein 65, putative isoform 3 | ||||
| STRING | EOY19496 | 0.0 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-147 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG044611t3 |
| Entrez Gene | 18586987 |



