 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG033777t1 | ||||||||
| Common Name | TCM_033777 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 360aa MW: 39276.4 Da PI: 10.1119 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 98.7 | 3.8e-31 | 287 | 344 | 3 | 59 |
-SS-EEEEEEE--TT-SS-EEEEEE-ST.T---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 3 DgynWrKYGqKevkgsefprsYYrCtsa.gCpvkkkversaedpkvveitYegeHnhe 59
D ++WrKYGqK++kgs++pr+YY+C+s gCp++k+ver+ +dp+++++tYeg+Hnh+
Thecc1EG033777t1 287 DDFSWRKYGQKPIKGSPHPRGYYKCSSVrGCPARKHVERAVDDPRMLIVTYEGDHNHS 344
99************************988****************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF10533 | 5.4E-18 | 238 | 283 | IPR018872 | Zn-cluster domain |
| Gene3D | G3DSA:2.20.25.80 | 8.2E-33 | 273 | 344 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 30.664 | 280 | 346 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 2.48E-26 | 283 | 344 | IPR003657 | WRKY domain |
| SMART | SM00774 | 5.4E-37 | 285 | 345 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 1.4E-26 | 287 | 343 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 360 aa Download sequence Send to blast |
MAVELMMGYG RGDSFAGKME ENALREAATA GIQGVEELIR LMSNSQQLYN QDASFKTSSP 60 SGPEPAMEIQ AVTDKTVNSF KKVISLLGRP RTGHARFRRA PLTSLQQEEK QQQQDLQQPR 120 QKIQESGTCS VQVNKDQVSA FKPFCPTPGH RLPPLPHNHH QSKSSPLLVA RSGLLERNEA 180 PTTINFTSSP PLSAGNSFIS SLTGDTDSMQ PSFSSGFQFT SPSHVPSSGK PPLSSSLKRK 240 CNSVDDAALK CGSASGRCHC SKKRKSRVKR VIRVPAISNK MADIPPDDFS WRKYGQKPIK 300 GSPHPRGYYK CSSVRGCPAR KHVERAVDDP RMLIVTYEGD HNHSHNITDA PAAVVLESS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 5e-21 | 273 | 343 | 2 | 71 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
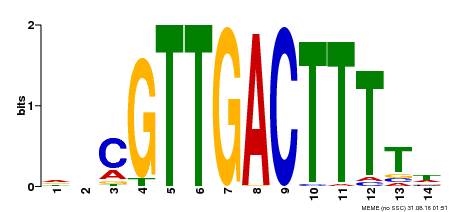
| UniProt | Transcription factor. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00450 | DAP | Transfer from AT4G24240 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salicylic acid and strongly during leaf senescence. {ECO:0000269|PubMed:11449049, ECO:0000269|PubMed:11722756}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007017155.2 | 0.0 | PREDICTED: probable WRKY transcription factor 7 | ||||
| Swissprot | Q9STX0 | 1e-120 | WRKY7_ARATH; Probable WRKY transcription factor 7 | ||||
| TrEMBL | A0A061FAK3 | 0.0 | A0A061FAK3_THECC; WRKY DNA-binding protein 7 | ||||
| STRING | EOY14380 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1715 | 28 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24240.1 | 1e-101 | WRKY DNA-binding protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG033777t1 |
| Entrez Gene | 18591144 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



