 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG031595t1 | ||||||||
| Common Name | TCM_031595 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 233aa MW: 26567.1 Da PI: 9.2858 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 138.2 | 2.4e-43 | 68 | 144 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
Cqve+C dls+ak+yhrrhkvCe+h+kap v+v+gl+qrfCqqCsrfhelsefDe+krsCrrrLa+hnerrrk++a
Thecc1EG031595t1 68 CQVEKCGLDLSDAKRYHRRHKVCEIHAKAPFVVVAGLRQRFCQQCSRFHELSEFDEAKRSCRRRLAGHNERRRKSSA 144
**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.6E-36 | 61 | 129 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 33.154 | 65 | 142 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.22E-40 | 66 | 145 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 2.7E-33 | 68 | 141 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010321 | Biological Process | regulation of vegetative phase change | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 233 aa Download sequence Send to blast |
METTDFRRKS ALKTKMNKGF IAEDLDDDME EEEEEGVGDH GFPDDEKKKK AYGKRGSSGG 60 AGVSPPSCQV EKCGLDLSDA KRYHRRHKVC EIHAKAPFVV VAGLRQRFCQ QCSRFHELSE 120 FDEAKRSCRR RLAGHNERRR KSSAESSTEG SSRKGMSSQL RESQCRQADD QRARVPIAIP 180 LSSVSQWFMH FTMYWYLLYY VGGFNNYIRH RNLSLVSVLL KPVFSCLACR VI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-37 | 56 | 141 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Promotes both vegetative phase change and flowering. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16914499}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00359 | DAP | Transfer from AT3G15270 | Download |
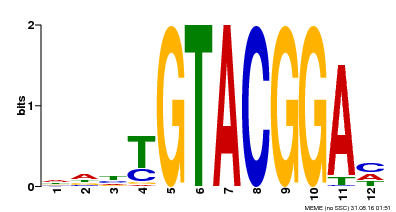
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156. {ECO:0000269|PubMed:16914499}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ622313 | 1e-133 | KJ622313.1 Gossypium hirsutum SQUAMOSA promoter binding-like transcription factor (SPL5) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017980011.1 | 1e-129 | PREDICTED: squamosa promoter-binding-like protein 4 | ||||
| Swissprot | Q9S758 | 2e-44 | SPL5_ARATH; Squamosa promoter-binding-like protein 5 | ||||
| TrEMBL | A0A061F852 | 1e-173 | A0A061F852_THECC; Squamosa promoter binding protein-like 3 isoform 1 | ||||
| STRING | EOY13069 | 1e-126 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G15270.1 | 4e-42 | squamosa promoter binding protein-like 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG031595t1 |
| Entrez Gene | 18594039 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



