 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG027630t2 | ||||||||
| Common Name | TCM_027630 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 483aa MW: 52779.8 Da PI: 6.1746 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 104.7 | 5.5e-33 | 257 | 311 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr+rWtpeLHe+Fveav++L G+ekAtPk +l+lm+v+gLt++hvkSHLQkYRl
Thecc1EG027630t2 257 KPRMRWTPELHECFVEAVSKLDGPEKATPKGVLKLMNVEGLTIYHVKSHLQKYRL 311
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.456 | 254 | 314 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 8.96E-18 | 255 | 311 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.2E-31 | 256 | 312 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-25 | 257 | 312 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.0E-9 | 259 | 310 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 2.4E-22 | 348 | 394 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 483 aa Download sequence Send to blast |
MKVGMSSKTM NHPSYISVTQ SEPSKGIGES HHTAASPIHN FLSIGSEGQS SLAGECSSPH 60 PFPFIRTESF KNNLKSGPSS PISPSSHAKS AFSRSSVFCT SLYLSSSSTS ETQRQLGNLP 120 FLPHPPTCGQ SISAVDSSKS PVVFSEDLHN PYNEDHSEII MKDFLNFPGD DCDGNFHGLH 180 CESNNFTLTE QLELQFLSDE LDIAIADHGE NPRLDEIYET PQKLNVAFTC NQNSASVVPS 240 TDACSSIRLS GPAAVHKPRM RWTPELHECF VEAVSKLDGP EKATPKGVLK LMNVEGLTIY 300 HVKSHLQKYR LAKYMPEKKE EKKTSSSEEK KAALSGNESD GKKKGGTHIT EALRMQMEVQ 360 KQLHEQLELQ RSLQLRIEEH ARYLQKILEE QQKAGSALLP SLSMSTPTDP SQNSELQPSS 420 SSAIASPTQP SESKTELSSS LPSKHKAPEV NDCEPESSPK KLRTENKPES AADEAVVENP 480 AQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 6e-26 | 257 | 314 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 6e-26 | 257 | 314 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 6e-26 | 257 | 314 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 6e-26 | 257 | 314 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 6e-26 | 257 | 314 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 6e-26 | 257 | 314 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 6e-26 | 257 | 314 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 6e-26 | 257 | 314 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
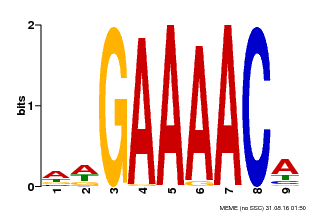
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007023582.2 | 0.0 | PREDICTED: myb family transcription factor PHL6 | ||||
| Refseq | XP_017979295.1 | 0.0 | PREDICTED: myb family transcription factor PHL6 | ||||
| Swissprot | Q949U2 | 1e-143 | PHL6_ARATH; Myb family transcription factor PHL6 | ||||
| TrEMBL | A0A061G9Y2 | 0.0 | A0A061G9Y2_THECC; Transcription factor, putative isoform 1 | ||||
| STRING | EOY26203 | 0.0 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 1e-104 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG027630t2 |
| Entrez Gene | 18595528 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



