 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG026767t1 | ||||||||
| Common Name | TCM_026767 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 375aa MW: 42028.1 Da PI: 4.8652 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 27.6 | 5e-09 | 305 | 338 | 22 | 55 |
SSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 22 nrypsaeereeLAkklgLterqVkvWFqNrRake 55
+yp++ee+ +L++ +gL+++q+ +WF N+R ++
Thecc1EG026767t1 305 WPYPTEEEKLQLSEITGLDQKQINNWFINQRKRH 338
59*****************************885 PP
| |||||||
| 2 | ELK | 36.8 | 8.5e-13 | 258 | 279 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
++K +L+rKYsgyL+sL++EF+
Thecc1EG026767t1 258 DIKGMLMRKYSGYLSSLRKEFL 279
58*******************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 4.7E-21 | 63 | 107 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 1.0E-20 | 65 | 106 | IPR005540 | KNOX1 |
| SMART | SM01256 | 8.2E-27 | 110 | 161 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 4.4E-25 | 113 | 160 | IPR005541 | KNOX2 |
| PROSITE profile | PS51213 | 10.58 | 258 | 278 | IPR005539 | ELK domain |
| SMART | SM01188 | 2.4E-6 | 258 | 279 | IPR005539 | ELK domain |
| Pfam | PF03789 | 1.3E-9 | 258 | 279 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.633 | 278 | 341 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.17E-19 | 280 | 347 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 6.3E-13 | 280 | 345 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-27 | 283 | 343 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.03E-12 | 290 | 342 | No hit | No description |
| Pfam | PF05920 | 6.0E-17 | 298 | 337 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 316 | 339 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 375 aa Download sequence Send to blast |
MEDMYRLDNP AISRSNDIVR VDNFAAANFS AAVTTDFLSP VDTILQFDHQ AADTDVTGSD 60 MSDLIKTQIA SHPRYPNLVS AYIECQKIGA PPELVSLLEE IGRENHPIRG CSEIGADPEL 120 DEFMESYCEV LHRYKEELSK PFDEATTFLS NIESQLSNLC KGALTKTLDY RSVNVVVEDS 180 NEFQSPATTV HSGRQLSRQT NFGLGFAYRL PFVHGEFCLT IKPKCSNEAA GTSEEELSGG 240 EVEASESQES AVARPSQDIK GMLMRKYSGY LSSLRKEFLK KRKKGKLPKD ARMTLLEWWN 300 NHYRWPYPTE EEKLQLSEIT GLDQKQINNW FINQRKRHWK PSEDMKFALF EGVAGNVGGP 360 VYLDAGVGTG SDNI* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 273 | 282 | LRKEFLKKRK |
| 2 | 279 | 283 | KKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during early embryogenesis. {ECO:0000269|PubMed:10488233}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00670 | PBM | Transfer from GRMZM2G087741 | Download |
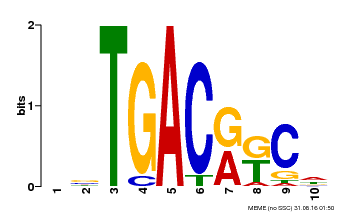
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX583168 | 3e-43 | JX583168.1 Gossypium hirsutum clone NBRI_GE16149 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021281551.1 | 0.0 | homeobox protein knotted-1-like 1 | ||||
| Swissprot | Q9FP29 | 4e-93 | KNOS1_ORYSJ; Homeobox protein knotted-1-like 1 | ||||
| TrEMBL | A0A061F2Z6 | 0.0 | A0A061F2Z6_THECC; KNOTTED1-like homeobox gene 6 isoform 1 | ||||
| STRING | EOY11655 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM835 | 27 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 2e-60 | KNOTTED-like from Arabidopsis thaliana | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG026767t1 |
| Entrez Gene | 18600575 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



