 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG021300t1 | ||||||||
| Common Name | TCM_021300 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 682aa MW: 74399.4 Da PI: 6.0876 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 91.3 | 8.3e-29 | 213 | 266 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Thecc1EG021300t1 213 KPRVVWSVELHQQFVAAVNQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 266
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 75.1 | 2.6e-25 | 35 | 143 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealk 91
vl+vdD+p+ +++l+++l+ y ev+ + +e al ll+e++ +D+++ D+ mp+mdG++ll++i e +lp+i+++a + + + + +
Thecc1EG021300t1 35 VLVVDDDPTCLMILEKMLRTCLY-EVTKCNRAETALSLLRENKngFDIVISDVHMPDMDGFKLLEHIGLEM-DLPVIMMSADDGKHVVMKGVT 125
89*********************.***************999999**********************6654.8******************** PP
TTESEEEESS--HHHHHH CS
Response_reg 92 aGakdflsKpfdpeelvk 109
Ga d+l Kp+ +e+l +
Thecc1EG021300t1 126 HGACDYLIKPVRIEALKN 143
**************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 3.1E-186 | 10 | 673 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 6.6E-42 | 32 | 172 | No hit | No description |
| SuperFamily | SSF52172 | 2.13E-35 | 32 | 156 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 2.6E-30 | 33 | 145 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.08 | 34 | 149 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 8.0E-23 | 35 | 143 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 2.80E-27 | 36 | 148 | No hit | No description |
| PROSITE profile | PS51294 | 11.048 | 210 | 269 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.58E-20 | 210 | 270 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-30 | 212 | 271 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.8E-25 | 213 | 266 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.8E-8 | 215 | 265 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0016310 | Biological Process | phosphorylation | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0016301 | Molecular Function | kinase activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 682 aa Download sequence Send to blast |
MNSSSGKGSM SAASSSAAWK AGDVVPDQFP AGLRVLVVDD DPTCLMILEK MLRTCLYEVT 60 KCNRAETALS LLRENKNGFD IVISDVHMPD MDGFKLLEHI GLEMDLPVIM MSADDGKHVV 120 MKGVTHGACD YLIKPVRIEA LKNIWQHVVR KRKNEWKDFE QSGSVEEGDR QPKQSEEADY 180 SSSANEGNWK SSKKRKDDDD EAEERDDTST LKKPRVVWSV ELHQQFVAAV NQLGIDKAVP 240 KKILELMNVP GLTRENVASH LQKYRLYLRR LSGVSQHQSN LNNSFMSPQE ATFGPLSPLN 300 GLDLQTLAAT GQLPAQSLAT FQAAGLGRST AKSGIAMPLV DQRNIFSFEN PKLRFGEGQQ 360 QHMNNNKQLN LLHGIPTTME PKQLASLHHS AQSIGNINMQ VTSHGVQGSQ NNSLLIQMAQ 420 PQPRGQILND STGSHAPRLP STLGQPILSN GIAANVSTRN GIPENIRGPG YNPVSQTSSL 480 LNFPMNHTSE LPGNSFPLGT TPGISSLTSK GAFQEDINSD VKGSGGFMPS YDIFNDLNQH 540 KPQNWELQNV GMTFDASQHS NSLQGNLDLA QSILVQQGFS SGQMNGQNRS AAVVSKAMFS 600 AGDCTEQGNA QNVNHHLNNL LVDNTIRIKS ERVADAGPAN LFPDHFGQED LMSALLKQQD 660 GIAPAENEFD FDGYSMDNIP V* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 4e-22 | 212 | 272 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
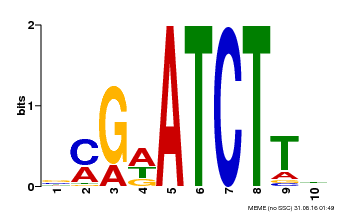
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007035713.2 | 0.0 | PREDICTED: two-component response regulator ARR2 | ||||
| Swissprot | Q9ZWJ9 | 0.0 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A061EWK1 | 0.0 | A0A061EWK1_THECC; Two-component response regulator | ||||
| STRING | EOY06639 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2403 | 27 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 0.0 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG021300t1 |
| Entrez Gene | 18603588 |



