 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG016909t1 | ||||||||
| Common Name | TCM_016909 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 473aa MW: 53418.8 Da PI: 6.9933 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 41.8 | 2.5e-13 | 235 | 271 | 11 | 48 |
HHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 11 llvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
llv++v+++G++ W++Ia++++ gR +kqc++rw+++l
Thecc1EG016909t1 235 LLVQLVTRHGTKKWSLIAKMLN-GRVGKQCRERWHNHL 271
79********************.*************97 PP
| |||||||
| 2 | Myb_DNA-binding | 54.1 | 3.6e-17 | 277 | 319 | 1 | 45 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
+++W+++Ed +l++a+k+ G++ W+ Iar+++ gRt++ +k++w+
Thecc1EG016909t1 277 KDSWSEDEDRILIEAHKEIGNK-WAEIARRLP-GRTENTIKNHWN 319
679*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 6.48E-5 | 205 | 259 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.738 | 206 | 271 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.0E-7 | 210 | 273 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-25 | 212 | 278 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.48E-12 | 214 | 271 | No hit | No description |
| Pfam | PF00249 | 1.9E-11 | 235 | 271 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 4.49E-27 | 250 | 329 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 26.038 | 272 | 326 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.3E-16 | 276 | 324 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.4E-15 | 277 | 319 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.05E-13 | 279 | 322 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-21 | 279 | 324 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010439 | Biological Process | regulation of glucosinolate biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1904095 | Biological Process | negative regulation of endosperm development | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 473 aa Download sequence Send to blast |
MEFETSLRED FPFLSNLFSD NSPLKPDFGN GFPLDGSSSS KGLLHNFILP DHNHYSPSVN 60 NGSLLNPYHF HQFPAEGSSR NPFFGFSSTC TDAFEPYVNG FSNDLNAYIP SLPFAPDGVN 120 NNGFLQGFQS EGCWDFSQKV PPQPLQPETQ TTYQPLIFQD QHGAVTAKVA DEVSCVTGDQ 180 NGYQDKVDQK KNKRFLTRRC SKAPKKSNII KGQWTPQEDS YIYEDLIMPI SFPRLLVQLV 240 TRHGTKKWSL IAKMLNGRVG KQCRERWHNH LRPDIKKDSW SEDEDRILIE AHKEIGNKWA 300 EIARRLPGRT ENTIKNHWNA TKRRQYSRRK GKDSNPKGAL LQSYIKSVSS TSTRKDKGKR 360 VMEANAQMLI NNNPAAKNPQ VVQVQSSDFN PKDWPVAVYN AQADHHQPMN FSFDASVFGE 420 SCTASFESML EEVPSGSLVE ESNAVMDFEL PLEMDSAKKE LDLLEMISQG NL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-43 | 208 | 325 | 4 | 107 | B-MYB |
| 1mse_C | 9e-44 | 209 | 325 | 2 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 9e-44 | 209 | 325 | 2 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 188 | 205 | QKKNKRFLTRRCSKAPKK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00385 | DAP | Transfer from AT3G27785 | Download |
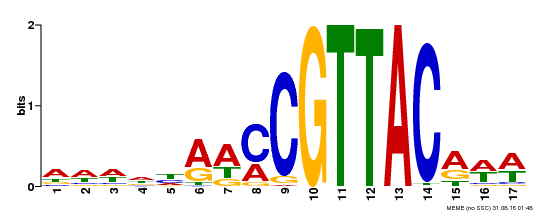
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017974434.1 | 0.0 | PREDICTED: transcription factor MYB98 | ||||
| TrEMBL | A0A061EBY4 | 0.0 | A0A061EBY4_THECC; Myb domain protein 118, putative | ||||
| STRING | EOY02441 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6052 | 26 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27785.1 | 2e-75 | myb domain protein 118 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG016909t1 |
| Entrez Gene | 18600785 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



