 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG016647t1 | ||||||||
| Common Name | TCM_016647 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 290aa MW: 32173.1 Da PI: 9.2314 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 104.3 | 7.2e-33 | 33 | 87 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprlrWt +LH+rFv+av++LGG++kAtPk++l+lm++kgLtl+h+kSHLQkYRl
Thecc1EG016647t1 33 KPRLRWTADLHDRFVDAVTKLGGPDKATPKSVLRLMGLKGLTLYHLKSHLQKYRL 87
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.455 | 30 | 90 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.3E-32 | 30 | 88 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.23E-16 | 33 | 90 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.0E-24 | 33 | 88 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 8.8E-9 | 35 | 86 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 1.8E-20 | 133 | 179 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 290 aa Download sequence Send to blast |
MERLSYGGGA GGGAVGGENF GYENGVVMTR DPKPRLRWTA DLHDRFVDAV TKLGGPDKAT 60 PKSVLRLMGL KGLTLYHLKS HLQKYRLGQQ ARKQNAVDQN KDNGGSSYVQ FSNHSPGTIT 120 NSPSADNDQR QIPVADALNT HLEVRRTLQE QLEVQKKLQM RVEAQGRYLQ AILEKAHKSL 180 SFDINCEGNV EETRAELTNF NLALSSLMEN VNGGADRKPN VVQMNEVPKK ATSCSAFQNY 240 AVGERERNKD VKLKVEGESI NFDLNTKDSF EFVAVNGNEL QSHMFSYKR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 8e-19 | 33 | 86 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 8e-19 | 33 | 86 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 8e-19 | 33 | 86 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 8e-19 | 33 | 86 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00544 | DAP | Transfer from AT5G45580 | Download |
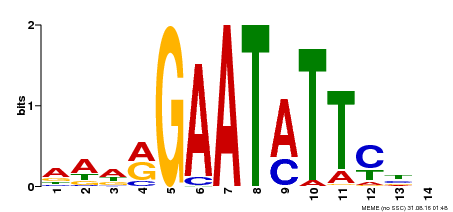
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX578082 | 2e-31 | JX578082.1 Gossypium hirsutum clone NBRI_GE10029 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007040781.2 | 0.0 | PREDICTED: myb family transcription factor PHL11 | ||||
| Swissprot | C0SVS4 | 4e-85 | PHLB_ARATH; Myb family transcription factor PHL11 | ||||
| TrEMBL | A0A061G803 | 0.0 | A0A061G803_THECC; Homeodomain-like superfamily protein, putative | ||||
| STRING | EOY25282 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM12495 | 25 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G45580.1 | 1e-74 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG016647t1 |
| Entrez Gene | 18606865 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



