 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG016510t1 | ||||||||
| Common Name | TCM_016510 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 985aa MW: 107729 Da PI: 4.9537 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 92 | 4.4e-29 | 577 | 627 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
Thecc1EG016510t1 577 EKSISLEVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 627
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.598 | 566 | 647 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 3.8E-26 | 580 | 627 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 1.83E-22 | 880 | 972 | No hit | No description |
| SMART | SM00666 | 8.2E-24 | 888 | 970 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 24.401 | 888 | 970 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 6.0E-17 | 889 | 969 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 4.7E-26 | 889 | 969 | No hit | No description |
| CDD | cd06407 | 3.27E-37 | 889 | 969 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 985 aa Download sequence Send to blast |
MCEPEEDNAC PSPPKQQQQL QGIMDLDDLD LESSWPLDQP TFLSNPTSPL IISSSSEQPC 60 SPLWAFSDED KVGSAAGYNL FLTCTPKPVN ENPKEDNDKR GIPSPFLGLL PLENPDSYCV 120 IKERMTQALR YFKDSTEQHV LAQVWAPIKS GGRYVLTTSG QPFVLDPHSN GLHQYRMVSL 180 MYMFSVDGES DGQLGLPGRV FRQKLPEWTP NVQYYSSKEY SRLDHALHYN VRGTLALPVF 240 EPSGQSCVGV LELIMTSQKI NYAPEVDKVC KALEAVNLKS SDILDPPSTQ ICNENRQNAL 300 AKILEILTVV CETYKLPLAQ TWVPCRHRSV LAYGGGLKKS CTSFDGSCMG QVCMSTTDVA 360 FYVVDAHMWG FREACLEHHL QKGQGVAGRA FLSRNSCFCT DITQFCKTEY PLVHYARMFR 420 LTSCFAICLR STYTGDDDYV LEFFLPPAIA DSNEQQTLLR SILATMKQHF QSLKVASGAE 480 LEDDEGSIEI IEASSDERLD SRLESIPIPP SVKSPPGPNT SPNRGELQLD SSKQQLIVTF 540 DPATDGGNVV ASGSQNPVCL PQNKDVKKSE RKRGKTEKSI SLEVLQQYFA GSLKDAAKSL 600 GVCPTTMKRI CRQHGISRWP SRKINKVNRS LTKLKHVIES VQGADGAFGL TSIATSPLPV 660 AVGSISWPTS LNGSNQQNSP NSKPSDPQGE KYDLPTCRTP VSNGQALVED QLLGGMTLSQ 720 EELFLQQNAL SPDLNKGANR SKTGSGSREE SAGTPTSHGS CQGSPAIESA ATKDPLSSIQ 780 EQCFKARGSP ELAFQPIGEL NIPATFSMPE ALVATEPQEP FGGMLVEDAG SSKDLRNLCP 840 SVADVGIDER FPESSWTPPP CTDLALMQAM ATFTQTTPHA TARQEMRSLT IKATYREDII 900 RFRISLSSGI VELKEEVAKR LKLEVGTFDI KYLDDDSEMV LIACDADLQE CLDVSRSSGS 960 NIIRLSVHDA MANLGSSCES TGEL* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
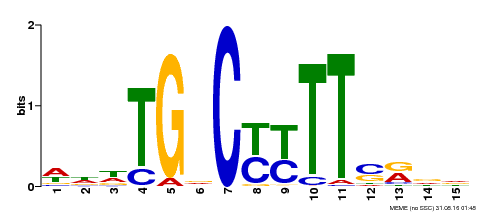
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017973247.1 | 0.0 | PREDICTED: protein NLP6 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A061G5G4 | 0.0 | A0A061G5G4_THECC; Transcription factor, putative | ||||
| STRING | EOY25090 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG016510t1 |
| Entrez Gene | 18606747 |



