 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG015714t1 | ||||||||
| Common Name | TCM_015714 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 670aa MW: 73148.9 Da PI: 5.5613 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.1 | 1.3e-12 | 490 | 535 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr ++P+ K++Ka+ L A+ YI++L
Thecc1EG015714t1 490 NHVEAERQRREKLNQRFYALRAVVPNV-----SKMDKASLLGDAISYINEL 535
799***********************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 3.5E-59 | 73 | 254 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.39 | 486 | 535 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 9.55E-19 | 489 | 560 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.85E-15 | 489 | 540 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 8.4E-19 | 490 | 556 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 4.4E-10 | 490 | 535 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 2.6E-16 | 492 | 541 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009269 | Biological Process | response to desiccation | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009867 | Biological Process | jasmonic acid mediated signaling pathway | ||||
| GO:0009963 | Biological Process | positive regulation of flavonoid biosynthetic process | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0043619 | Biological Process | regulation of transcription from RNA polymerase II promoter in response to oxidative stress | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051090 | Biological Process | regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:2000068 | Biological Process | regulation of defense response to insect | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 670 aa Download sequence Send to blast |
MTDYRLATAI NLWTDDNASV MEAFMSSDLS ALWPPPQSSG STSAPAAAAG PDPSKSSLAQ 60 SQPSVSLLNQ ETLQQRLQAL IEGARENWTY AIFWQSSYDY SGTAVLGWGD GYYKGEEDKG 120 KGKLKASSST AAEQEHRKKV LRELNSLISG STSPTDDAVD EEVTDTEWFF LVSMTQSFVN 180 GGGLPGQAFF NSSPVWVAGS DRLATSICER ARQGQVFGLQ TMVCIPSANG VVELGSTELI 240 TQSSDLMNKV RVLFNFNNGI EAGSWSMSNN TADQGENDPS SLWINDPNNG IELKESNNNS 300 NNNNTSHQNQ QIQKSIQFCD NPSSSSLTEN PSSIHVGNHQ QQQNHQQGHS FCLNFSDYGF 360 DGSSSVRNGN SSSHLLKPES GEILNFGESK RSGNGNLFSG NSQIGVEENK KKRSPTSRGS 420 NEEGMLSFTS GVILPSSGVV KSSGGAGDSD HSDLEASVVK EADSSRVVEP EKRPRKRGRK 480 PANGREEPLN HVEAERQRRE KLNQRFYALR AVVPNVSKMD KASLLGDAIS YINELRTKLQ 540 NADSEKEELQ KELEAMKKEL SSKDSRSAPP APDQDLKMSN HLGNKLVELE IDVKIIGWDA 600 MIRIQCNKKN HPAARLMAAL KELDLDVHHA SVSVVNDLMI QQATVKMGSR FYTQEQLRIA 660 LTSKFGDAR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 1e-79 | 68 | 257 | 4 | 195 | Transcription factor MYC3 |
| 4rqw_B | 1e-79 | 68 | 257 | 4 | 195 | Transcription factor MYC3 |
| 4rs9_A | 1e-79 | 68 | 257 | 4 | 195 | Transcription factor MYC3 |
| 4yz6_A | 1e-79 | 68 | 257 | 4 | 195 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 471 | 479 | KRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds to the G-box motif (5'-AACGTG-3') found in the promoter of the jasmonate-induced gene LAPA1 (PubMed:15231736). Acts as negative regulator of blue light-mediated photomorphogenesis and positively regulates root growth (PubMed:24483714). Promotes growth in response to the phytohormones abscisic acid (ABA) and jasmonate (JA) (PubMed:24483714). Binds to the G-box motif (5'-CACGTG-3') of the RBCS-3A gene promoter (PubMed:24483714). Acts downstream of the jasmonate (JA) receptor to orchestrate JA-mediated activation of plant responses (PubMed:28733419). Positively regulates both wound-responsive and pathogen-responsive genes through MYC2-targeted transcription factors (MTFs) involved in early response to JA (PubMed:28733419). With JA2L forms a transcription module that regulates wounding-responsive genes (PubMed:28733419). With ERF.C3 forms a transcription module that regulates pathogen-responsive genes (PubMed:28733419). Plays a critical role in orchestrating JA-mediated defense gene expression during Botrytis cinerea infection (PubMed:28733419). Regulates negatively defense responses to root-knot nematodes, potentially by mediating crosstalk among the hormones strigolactones, abscisic acid (ABA) and jasmonate (JA) (PubMed:30576511). Regulates the termination of JA-mediated defense responses by specifically binding the G-box (5'-CACATG-3') motifs in the promoters of MTB1, MTB2 and MTB3, which are transcription factors that negatively regulates JA signaling (PubMed:30610166). May be involved in JA-induced chilling tolerance, possibly by ameliorating the antioxidant enzyme system of fruit and increasing proline and lycopene levels (PubMed:29528226). {ECO:0000269|PubMed:15231736, ECO:0000269|PubMed:24483714, ECO:0000269|PubMed:28733419, ECO:0000269|PubMed:29528226, ECO:0000269|PubMed:30576511, ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00084 | PBM | Transfer from AT1G32640 | Download |
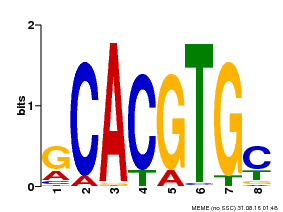
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by methyl jasmonate (PubMed:15231736, PubMed:28733419). Induced by wounding (PubMed:15231736). Induced by infection with the fungal pathogen Botrytis cinerea (PubMed:28733419). Induced in fruit by storage in cold (PubMed:29528226). Induced by hydrogen peroxide and infection with root-knot nematodes (PubMed:30576511). {ECO:0000269|PubMed:15231736, ECO:0000269|PubMed:28733419, ECO:0000269|PubMed:29528226, ECO:0000269|PubMed:30576511}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017973469.1 | 0.0 | PREDICTED: transcription factor MYC2 | ||||
| Swissprot | A0A3Q7HRZ6 | 0.0 | MYC2_SOLLC; Transcription factor MYC2 | ||||
| TrEMBL | A0A061GAC4 | 0.0 | A0A061GAC4_THECC; Basic helix-loop-helix DNA-binding family protein | ||||
| STRING | EOY23994 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1265 | 28 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G17880.1 | 2e-94 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG015714t1 |
| Entrez Gene | 18606039 |



