 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG014549t4 | ||||||||
| Common Name | TCM_014549 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1197aa MW: 136159 Da PI: 8.1287 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.3 | 0.00026 | 1089 | 1111 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
Thecc1EG014549t4 1089 CPvkGCGKKFFSHKYLVQHRRVH 1111
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 12.1 | 0.0006 | 1147 | 1173 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
Thecc1EG014549t4 1147 YVCAeeGCGQTFRFVSDFSRHKRKtgH 1173
89********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 26 | 1064 | 1086 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.362 | 1087 | 1116 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0045 | 1087 | 1111 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.6E-5 | 1088 | 1115 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1089 | 1111 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.6E-9 | 1103 | 1143 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.7E-8 | 1116 | 1141 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 1117 | 1146 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1117 | 1141 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1119 | 1141 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.32E-8 | 1135 | 1169 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-9 | 1142 | 1170 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.24 | 1147 | 1178 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.85 | 1147 | 1173 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1149 | 1173 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1197 aa Download sequence Send to blast |
MSRGLCNYKD VVKLSKDLAS DEIMVGGNEE IKGVKGFYSV KGKFASMYEG NRDSAFNGTD 60 HLCRLPLQTL NMSAEGENAV QGDALSDQGL FSCVTCGILC FSCIAVLQPT EQAARYLMSA 120 DCSFFNDWTV GSGVTRDGFT TTHGDVITSE QNSCTRWMNK RAPNALYDVP VQSVEDKFHM 180 ADQSNQVVED TEKGGDTSAL GLLASTYGNS SDSEEDHVEP NVTVSGDETN SANRSLERKF 240 QYNGSGFSPG DANGSNNPSL LRLESEEEAP VHVDIKSTSP QAFDHTVEFE TDNLASRRSI 300 GLEDKFRDPI TTSHANPSYS PATHGAEKMR FSKTMVPMEN ADIPFAPRSD EDSSRMHVFC 360 LEHAVEVDQQ LRQIGGVHVF LLCHPEYPKI EAEAKLVTEE LGIDYPWNDI LFGDATKEDE 420 ERIQSALDSE DAIPGNGDWA VKLGVNLFYS ANLSRSTLYS KQMPYNYVIY SAFGRNSPGS 480 SPTKLNVYGR RSGKQKKVVA GKWCGKVWMS NQVHPFLAQR DPEEQEQERG FHAWATSDEN 540 LERKPENVHK AETTKVAKKF NRKRKMRPEI ASSKKVKCIE TEGAVSDDSL DGGSLRQQQI 600 FFRGKQPRLI QKEEAISYDL LEDDSLLQQR NLSRKKLAKF IEREGAESED AEEEFTHQQH 660 WRNLRGKQGK YIEEDDAVSG DSLDESSLKQ YRRIPRSWQA KFREREDIVS EDELEEISHR 720 LHRRIPRCRQ IKSCEKNDAI SDDSRADNSL KQYRRMPKGR QANFVERDDT MSDDASEDDS 780 QHQLRRIPKG KQMKCMERDD AFSDDSLEDN LQQQHRIPRS KVAKFTDRED VVSFDSLKGS 840 SHQQRRRVSR SQLTKFIERE DAVSSDSPDD SSLQQLRRIP RSKQTKILER EDAVSDDSLD 900 DTSQQQLRKT PRSRQGKFIE REDAVSYDSL DENYHQPNRR TLRSRKKKAQ TPRQIKQETP 960 RNVKQGKRRT TKQVVSQQIK QETPRNRNTK IEQSARQCNS YGEDELEGGP STRLRKRVRK 1020 PLKESETKPK EKKQASKKKV KNASNVKTLA GHNTSKVRDE EAEYQCDMEG CTMSFGLKQE 1080 LLLHKRNICP VKGCGKKFFS HKYLVQHRRV HLDDRPLKCP WKGCKMTFKW AWARTEHIRV 1140 HTGARPYVCA EEGCGQTFRF VSDFSRHKRK TGHSAKKGLG STKGLEFPHC CHSGNC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 6e-69 | 1062 | 1177 | 21 | 136 | Lysine-specific demethylase REF6 |
| 6a58_A | 6e-69 | 1062 | 1177 | 21 | 136 | Lysine-specific demethylase REF6 |
| 6a59_A | 6e-69 | 1062 | 1177 | 21 | 136 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1012 | 1018 | RLRKRVR |
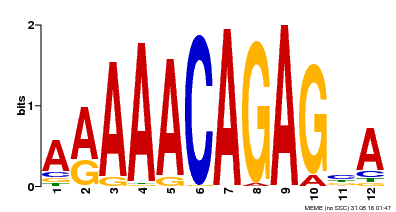
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007037855.2 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A061G5N9 | 0.0 | A0A061G5N9_THECC; Relative of early flowering 6, putative isoform 4 | ||||
| STRING | EOY22356 | 0.0 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 1e-87 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG014549t4 |
| Entrez Gene | 18605027 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



