 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG013043t10 | ||||||||
| Common Name | TCM_013043 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 764aa MW: 88028.6 Da PI: 8.9354 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 85.6 | 7.4e-27 | 11 | 113 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgkwevtk 80
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ +++ +r+ ++t+Cka+++vk++ dgkw+v++
Thecc1EG013043t10 11 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSfnrprarqskqdPDNTTGRRSCSKTDCKASMHVKRRPDGKWVVHS 102
5*******************************************9999999987766554444558999*********************** PP
FAR1 81 leleHnHelap 91
+++eHnHel p
Thecc1EG013043t10 103 FVKEHNHELLP 113
********975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.2E-24 | 11 | 113 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 4.7E-29 | 211 | 303 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.545 | 491 | 527 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 5.3E-4 | 501 | 524 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.8E-7 | 502 | 529 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 764 aa Download sequence Send to blast |
MEFESHGEAY SFYQEYARSM GFNTAIQNSR RSKTSREFID AKFACSRYGT KREYDKSFNR 60 PRARQSKQDP DNTTGRRSCS KTDCKASMHV KRRPDGKWVV HSFVKEHNHE LLPAQAVSEQ 120 TRRMYAAMAR QFAEYKNVVG LKNDPKNPFD KGRNLALEAG DVKILLEFFT HMQNINSNFF 180 YAIDLGEDQR LKSLFWVDAK SRHDYSYFCD VVSFDTTYVR NKYKMPLALF IGVNHHYQFM 240 PLGCALVSDD SAATFSWLMQ TWLKAMGGQS PRVIITDQDR IVKSVVAEIF PNTHHCFFLW 300 HVLGKVSENL GHVIKQHGNF MAKFEKCIYR SWTEEEFAKR WWKILDRFGL KDDEWMKSLY 360 EDRRKWVPTY IMDVLLAGMS MVQRSESVNS FFDKYVHKKT TVQEFLKQYE AILQDRYEEE 420 AKANSDSWSK LPTLKSPSPF EKSVAGLYTH TVFKKFQVEV VGAIACHPKP ENHDATSSFF 480 RVQDLEKNQD FIVTLNEMKS EVSCICRLYE YKGYLCRHAM VVLQINGHSA IPSQYILKRW 540 TKEAKSRHLM GDESEQVQSR VQRYNDLFQR AMKLIEEGSL SQESYYIAFR SLEEAFGNCL 600 SANTSNKSLA EAVTSPTQGM ICIEEDNQSR STSKTNKKKN PTKKRKGNSE QEVMTVPATD 660 GLQQMDKLSS RSVGLDGYFG AQTSVQGMLN LMAPRDNYYG NQQTIQGLGQ LNTIAASHDG 720 YYGPQQTMPG MGQMDFFRAP GFYIRDDTNV RAAQLHDDAS RHA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
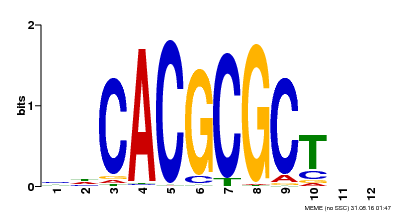
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017973539.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X2 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A061FW45 | 0.0 | A0A061FW45_THECC; Far-red elongated hypocotyls 3 isoform 10 | ||||
| STRING | EOY21471 | 0.0 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG013043t10 |
| Entrez Gene | 18604429 |



