 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG011862t1 | ||||||||
| Common Name | TCM_011862 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 359aa MW: 39939.1 Da PI: 8.8008 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 44 | 5.6e-14 | 63 | 111 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f ++eAaka++ a+++++g
Thecc1EG011862t1 63 SKYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEDEAAKAYDIAAQRFRG 111
89****9888.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 100 | 1.4e-31 | 192 | 289 | 1 | 93 |
EEEE-..-HHHHTT-EE--HHH.HTT.....---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.....ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f+k+ tpsdv+k++rlv+pk++ae++ g+ + +++ l +ed +g++W+++++y+++s++yvltkGW++Fvk+++Lk+gD+v F+++
Thecc1EG011862t1 192 FEKAVTPSDVGKLNRLVIPKQHAEKYfplqsGSASSKGVLLNFEDVTGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKNLKAGDIVSFQRSTGL 284
89****************************97788899**************************************************76554 PP
EE..E CS
B3 89 efelv 93
e++l+
Thecc1EG011862t1 285 EKQLY 289
44455 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 4.71E-17 | 63 | 120 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 8.5E-9 | 63 | 111 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 7.83E-26 | 63 | 121 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 4.6E-20 | 64 | 120 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.126 | 64 | 119 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.4E-29 | 64 | 125 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 1.8E-42 | 186 | 296 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.19E-31 | 190 | 295 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 7.37E-28 | 191 | 281 | No hit | No description |
| PROSITE profile | PS50863 | 14.311 | 192 | 294 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 6.8E-29 | 192 | 292 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.8E-24 | 192 | 296 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 359 aa Download sequence Send to blast |
MDGSCIDEST TSGDSISISP SNLSPLPPAT KSPDSLCRVG SGASVIVDSE AGVEAESRKL 60 PSSKYKGVVP QPNGRWGAQI YEKHQRVWLG TFNEEDEAAK AYDIAAQRFR GRDAVTNFKH 120 LRETEEDDIE MAFLNSHSKA EIVDMLRKHT YNDELEQSRR SYGFDGNGKR VVRHDGGFGS 180 FGLELKAREQ LFEKAVTPSD VGKLNRLVIP KQHAEKYFPL QSGSASSKGV LLNFEDVTGK 240 VWRFRYSYWN SSQSYVLTKG WSRFVKEKNL KAGDIVSFQR STGLEKQLYI DWKTRTGLGS 300 GLENPVGPVQ MVRLFGVNIF KIPGSENVGI VGCIGKRTRE MELLELECSK KQRVIDAL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 5e-60 | 189 | 297 | 11 | 119 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds specifically to bipartite recognition sequences composed of two unrelated motifs, 5'-CAACA-3' and 5'-CACCTG-3'. May function as negative regulator of plant growth and development. {ECO:0000269|PubMed:15040885, ECO:0000269|PubMed:9862967}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
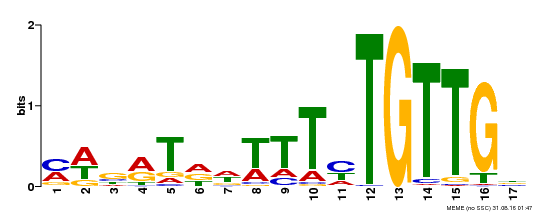
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by brassinosteroid and zeatin. {ECO:0000269|PubMed:15040885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ801819 | 0.0 | KJ801819.1 Gossypium hirsutum RAV protein (RAV1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007046304.2 | 0.0 | PREDICTED: AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| Swissprot | Q9ZWM9 | 1e-156 | RAV1_ARATH; AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| TrEMBL | A0A061EB30 | 0.0 | A0A061EB30_THECC; AP2 domain class transcription factor | ||||
| STRING | EOY02136 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM848 | 27 | 121 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68840.2 | 1e-151 | related to ABI3/VP1 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG011862t1 |
| Entrez Gene | 18610537 |



