 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG008193t1 | ||||||||
| Common Name | TCM_008193 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 427aa MW: 46915.8 Da PI: 6.4402 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 88 | 9.2e-28 | 153 | 207 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k +++WtpeLH+rFv+aveqL G +kA+P++ilelm++++Lt+++++SHLQkYR++
Thecc1EG008193t1 153 KVKVDWTPELHRRFVQAVEQL-GVDKAVPSRILELMGIDCLTRHNIASHLQKYRSH 207
5689*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 18.152 | 150 | 209 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.23E-18 | 151 | 210 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-26 | 151 | 210 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.8E-26 | 153 | 207 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.5E-8 | 156 | 205 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0010638 | Biological Process | positive regulation of organelle organization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 427 aa Download sequence Send to blast |
MLAVSPLRNT TNDENKGEME SFTISSEEFP DFADGNLLES IDFDDLFVSI NEGDMLPDLE 60 MDPEIIAELP TNGSEESEMN TSIDKTDEDN NQRKEEEDKV SGSGSGLGSS SSKGEEIVSK 120 REEPTAVKTP TKDADKGRKS SLQAKNNNQG KRKVKVDWTP ELHRRFVQAV EQLGVDKAVP 180 SRILELMGID CLTRHNIASH LQKYRSHRKH LLAREAEAAS WTHRRQMYGA ATPAGGGKRD 240 MNPWLAPTMG FPPMSPMHHH HHFRPLHVWG HPTVDQSLMH LWPKHLAHTP SPSPPPPPTW 300 GPHPPPADPS YWHHHDQRVP NGLTPGTPCF PQPLAPTRFA APPVPGIPPH HPMYKADPGI 360 GVPAGQSGPH PLIDFHPSKE SIDAAIEDVL SKPWLPLPLG LKPPSTDSVL GELQRQGVPK 420 VPPSCA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 1e-15 | 150 | 205 | 2 | 57 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that functions with GLK2 to promote chloroplast development. Acts as an activator of nuclear photosynthetic genes involved in chlorophyll biosynthesis, light harvesting, and electron transport. Acts in a cell-autonomous manner to coordinate and maintain the photosynthetic apparatus within individual cells. May function in photosynthetic capacity optimization by integrating responses to variable environmental and endogenous cues (PubMed:11828027, PubMed:12220263, PubMed:17533111, PubMed:18643989, PubMed:19376934, PubMed:19383092, PubMed:19726569). Prevents premature senescence (PubMed:23459204). {ECO:0000269|PubMed:11828027, ECO:0000269|PubMed:12220263, ECO:0000269|PubMed:17533111, ECO:0000269|PubMed:18643989, ECO:0000269|PubMed:19376934, ECO:0000269|PubMed:19383092, ECO:0000269|PubMed:19726569, ECO:0000269|PubMed:23459204}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
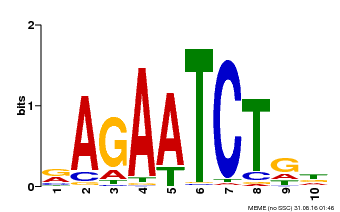
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. Repressed by BZR2. {ECO:0000269|PubMed:12220263, ECO:0000269|PubMed:21214652}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021279775.1 | 0.0 | transcription activator GLK1 | ||||
| Swissprot | Q9SIV3 | 1e-113 | GLK1_ARATH; Transcription activator GLK1 | ||||
| TrEMBL | A0A061E3A4 | 0.0 | A0A061E3A4_THECC; GBF's pro-rich region-interacting factor 1 | ||||
| STRING | EOX99519 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2209 | 27 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 3e-97 | GBF's pro-rich region-interacting factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG008193t1 |
| Entrez Gene | 18608769 |



