 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG007074t1 | ||||||||
| Common Name | TCM_007074 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 424aa MW: 46168.9 Da PI: 9.8149 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 45.4 | 1.8e-14 | 347 | 399 | 5 | 57 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkeva 57
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++ N++L+k+ ee+ ++ +
Thecc1EG007074t1 347 RRQRRMIKNRESAARSRARKQAYTLELEAEVAKLKEMNQELQKKQEEMMEMQK 399
79*****************************************9999998865 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.2E-12 | 343 | 407 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.875 | 345 | 396 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 8.1E-14 | 347 | 397 | No hit | No description |
| Pfam | PF00170 | 1.2E-12 | 347 | 399 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 1.03E-25 | 347 | 401 | No hit | No description |
| SuperFamily | SSF57959 | 4.56E-10 | 347 | 397 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 350 | 365 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 424 aa Download sequence Send to blast |
MGSHLNYKNF GDAPSMEGNG SKPLGNFPLA RQSSIYSLTF DELQNTFGGL GKDFGSMNMD 60 ELLKNISTAE ETQAVTTASV AGGEGSFSGG NLQRQGSLTL PRTLSQKTVD EVWRDLMKEN 120 DGAKDGSNGG GVGGGASLPQ RQQTLGEMTL EEFLVRAGVV REDMQQIGMA NNSGFFGNNT 180 GVALGFQQTN RNNGFLSNNN SVLNHPPRLP LNMSGVKSSQ PQQQQQQQQQ QQPQQQQQQQ 240 PQPLFPKQQT VAFAPSMHLM NTAQLANPGV RGSMVGIGDP SMNSNLVQST GLQSGGMGIV 300 GLASPPSQIS SDMIPNNSVD ATSLSPVPYV FGRGRKCSAA LEKVVERRQR RMIKNRESAA 360 RSRARKQAYT LELEAEVAKL KEMNQELQKK QEEMMEMQKN QMLETMSRPC KRHCLRRTLT 420 GPW* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
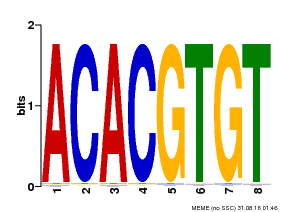
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007042440.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Swissprot | Q9M7Q3 | 1e-127 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | A0A061DZZ4 | 0.0 | A0A061DZZ4_THECC; Abscisic acid responsive element-binding factor 1, putative isoform 1 | ||||
| STRING | EOX98270 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM734 | 27 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49720.1 | 7e-89 | abscisic acid responsive element-binding factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG007074t1 |
| Entrez Gene | 18607945 |



