 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG005705t1 | ||||||||
| Common Name | TCM_005705 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 271aa MW: 29946.3 Da PI: 4.8543 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 62.8 | 7.7e-20 | 133 | 184 | 2 | 56 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkklege 56
++kGVr+++ +g+++AeIrdp+ ng +r++lg++ t+e+Aa a+++a+ k++g+
Thecc1EG005705t1 133 HFKGVRRRP-WGKYAAEIRDPKRNG--ARIWLGTYETPEDAALAYDRAAFKMRGA 184
58*******.**********98776..**************************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 5.6E-14 | 133 | 183 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.327 | 133 | 191 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.7E-31 | 133 | 192 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.98E-30 | 133 | 193 | No hit | No description |
| SuperFamily | SSF54171 | 3.4E-23 | 133 | 192 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 2.3E-37 | 133 | 197 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.3E-12 | 134 | 145 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.3E-12 | 173 | 193 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 271 aa Download sequence Send to blast |
MNVEIMSGSE FNILESFQCY LLDDFEESQA AAATTTTSNL VVLEPSYNRS SSFSGVFLNE 60 SWGDLPLKID DSEDMLVYSA LRDAANSGWS PSNEVDNYTP VAVEELVSTT TREIEKEESK 120 VEPETSVPPS GMHFKGVRRR PWGKYAAEIR DPKRNGARIW LGTYETPEDA ALAYDRAAFK 180 MRGAKAKLNF PHLIGSNTWE PIRVGGRRRS PESSAPTTAS SLSDCSRKRR KSSASNSDQV 240 TAKLREGASL AAEVIQMTTP WTLGDQFWII * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 3e-30 | 131 | 197 | 3 | 69 | ATERF1 |
| 3gcc_A | 3e-30 | 131 | 197 | 3 | 69 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. Involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways. {ECO:0000269|PubMed:11950980}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00311 | DAP | Transfer from AT2G44840 | Download |
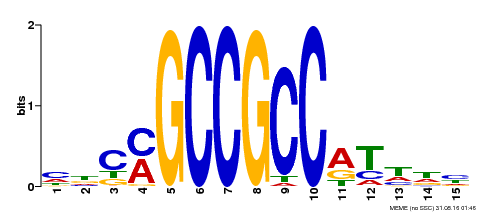
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Strongly induced by wounding. Induced by Pseudomonas syringae tomato (both virulent and avirulent avrRpt2 strains), independently of PAD4. Also induced by methyl jasmonate (MeJA) independently of JAR1, but seems to not be affected by ethylene. Induction by salicylic acid (SA) is controlled by growth and/or developmental conditions, and seems to be more efficient and independent of PAD4 in older plants. {ECO:0000269|PubMed:11950980, ECO:0000269|PubMed:12068110}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007052302.2 | 0.0 | PREDICTED: ethylene-responsive transcription factor 13 | ||||
| Swissprot | Q8L9K1 | 9e-65 | ERF99_ARATH; Ethylene-responsive transcription factor 13 | ||||
| TrEMBL | A0A061DUI1 | 0.0 | A0A061DUI1_THECC; Ethylene-responsive element binding factor 13, putative | ||||
| STRING | EOX96459 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10 | 28 | 1650 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G44840.1 | 1e-63 | ethylene-responsive element binding factor 13 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG005705t1 |
| Entrez Gene | 18614468 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



