 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG004922t1 | ||||||||
| Common Name | TCM_004922 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 982aa MW: 108240 Da PI: 6.4526 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 128.1 | 3.3e-40 | 149 | 226 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls k+yhrrhkvCe+hska+++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Thecc1EG004922t1 149 VCQVEDCGADLSCSKDYHRRHKVCEMHSKASKALVGNVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTNP 226
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 7.3E-33 | 142 | 211 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.829 | 147 | 224 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.22E-37 | 148 | 230 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 6.0E-29 | 150 | 223 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 1.4E-5 | 752 | 877 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.35E-6 | 752 | 864 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.63E-6 | 753 | 864 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 982 aa Download sequence Send to blast |
MEARFGSDAH HFYGMNPANL RAVGKRTLEW DLNDWKWDGD LFIASSINPV SADSTGRQFF 60 PLGSGIPGNS SNSSSSCSDE VNLETEKGKR ELEKKRRVIV VEDDSPNEEA GSLTLKLGGQ 120 GGHGYPISQR EGTSGKKTKL GGGSGNRAVC QVEDCGADLS CSKDYHRRHK VCEMHSKASK 180 ALVGNVMQRF CQQCSRFHVL QEFDEGKRSC RRRLAGHNKR RRKTNPDTVV NGNSLNDEQT 240 SGYLLLSLLK ILSNMHSNRS DQTTDQDVLS HLLRSLANHT GEQGGRNISG LLPEPQDSEA 300 VSALFLNGQG PPRPFKQHHT GAASEMAEKG VSSQGTRGVK VQGNTAGAVK MNNFDLNDIY 360 IDSDEGTDDI ERSPAAVNTG TSSLDCPSWI QQDSHQSSPP QTSGNSDSAS AQSPSSSSGD 420 AQSRTDRIVF KLFGKEPNDF PMVLRAQILD WLSHSPTDIE SYIRPGCIVL TIYLRQAEAA 480 WDELCCDLSF TLSRLLDCSD DTFWRSGWIY IRVQDQIAFI YNGQVVVDTS LPLRSNHYSK 540 ITSVKPIAIS ATERAQFSVK GINLSRPATR LLCAVEGKCL LQETTNELMD GNDDYKEQDE 600 LQCVNFSCSV PTVTGRGFIE IEDHGFSSSF FPFIVAEEDV CSEVRMLESV LEISDTDADV 660 GGTGKLEAKH RAMDFIHEVG WLLHRCQLKS RLGHLDPNPE PFPLSRFKWL MEFSMDHEWC 720 AVVKKLLNIL LNGVVGSGEH PSLNLALTEM GLLHRAVRKN CRPLVELLLR FVPEKASDKL 780 GFENETLTGV DHKSFLFRPD VLGPAGLTPL HIAAGKDGSE DVLDALTDDP GKVGIDAWKS 840 ARDSTGSTPE DYARLRGHYS YIHLVQKKIN KRTASGHVVV DIPGALSECS MNQKQNNEST 900 SSFEIGRLEL RSIQRHCKLC DQKLAYGCGT TSKSLVYRPA MLSMVAIAAV CVCVALLFKS 960 CPEVLYVFRP FRWELLDYGT S* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 6e-30 | 142 | 223 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
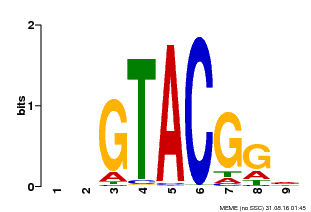
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ622325 | 0.0 | KJ622325.1 Gossypium hirsutum SQUAMOSA promoter binding-like transcription factor (SPL17) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017979528.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A061DRH1 | 0.0 | A0A061DRH1_THECC; Squamosa promoter-binding protein, putative isoform 1 | ||||
| STRING | EOX95415 | 0.0 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG004922t1 |
| Entrez Gene | 18613790 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



