 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG002318t1 | ||||||||
| Common Name | TCM_002318 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 504aa MW: 54950.9 Da PI: 6.5998 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 97.9 | 6.4e-31 | 107 | 164 | 2 | 60 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS-- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhek 60
+DgynWrKYGqK vkg+ef rsYY+Ct+++C vkk++ers+ d+k+v+++Y g+H+h+k
Thecc1EG002318t1 107 EDGYNWRKYGQKLVKGNEFVRSYYKCTHPNCLVKKQLERSH-DGKMVDTVYFGQHDHPK 164
7****************************************.***************85 PP
| |||||||
| 2 | WRKY | 103.4 | 1.2e-32 | 281 | 339 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
++Dgy+WrKYGqK vkg+++prsYYrC+ +gCpvkk+ver+++d k+v++tYeg+H+h+
Thecc1EG002318t1 281 VNDGYRWRKYGQKLVKGNPNPRSYYRCSNPGCPVKKHVERDSHDVKLVITTYEGRHDHD 339
58********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50811 | 23.824 | 101 | 165 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 7.6E-27 | 102 | 165 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.12E-24 | 103 | 165 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.7E-35 | 106 | 164 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 1.1E-23 | 107 | 163 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 5.4E-35 | 267 | 341 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.22E-28 | 273 | 341 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 35.714 | 276 | 341 | IPR003657 | WRKY domain |
| SMART | SM00774 | 2.2E-37 | 281 | 340 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 2.3E-25 | 282 | 338 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009863 | Biological Process | salicylic acid mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 504 aa Download sequence Send to blast |
MVSSGECITD EVGTDKLQSP DAGNHALQQI SDSKVHLPQS NQGGETPSIK SEKAPLPDSM 60 VQTSKSEQGG NVSCMISNKT SVTPDMTPSP LPGTEGSTPI VREKASEDGY NWRKYGQKLV 120 KGNEFVRSYY KCTHPNCLVK KQLERSHDGK MVDTVYFGQH DHPKPLNLPV AVGVVVSVVE 180 EKPYNASPTV VKDKSSDARS QTPCQIEPQD DFRPLAIAAS ENVKGAPSKS IRIQNVADSD 240 DDHVISKRRK KENSNADASP VEKQTTESRM VVKTLSEVDI VNDGYRWRKY GQKLVKGNPN 300 PRSYYRCSNP GCPVKKHVER DSHDVKLVIT TYEGRHDHDI PPTRTVTHNT TGVNVHSAAH 360 NDESGTKVEE SETACVDMVV HSSSGLENKS SEQLNGESTT KSEASGAVCV DVVEAPISGP 420 ESGSNEQRSG KLQPSKRSKG DGNDMIVHSK SISQNTAKEQ LSNKLETKSE NDTVCIDKMV 480 HITPHPECNF DEQRMPSAEP VQS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 1e-37 | 108 | 342 | 1 | 75 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Binds to a 5'-CGTTGACCGAG-3' consensus core sequence which contains a W box, a frequently occurring elicitor-responsive cis-acting element. {ECO:0000269|PubMed:8972846}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00094 | SELEX | Transfer from AT2G04880 | Download |
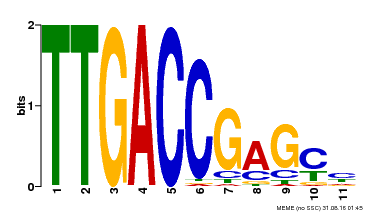
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salicylic acid (SA). {ECO:0000269|PubMed:17264121}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007049282.2 | 0.0 | PREDICTED: WRKY transcription factor 1 | ||||
| Refseq | XP_007049283.2 | 0.0 | PREDICTED: WRKY transcription factor 1 | ||||
| Refseq | XP_007049284.2 | 0.0 | PREDICTED: WRKY transcription factor 1 | ||||
| Swissprot | Q9SI37 | 3e-95 | WRKY1_ARATH; WRKY transcription factor 1 | ||||
| TrEMBL | A0A061DMX7 | 0.0 | A0A061DMX7_THECC; Zinc-dependent activator protein-1, putative isoform 1 | ||||
| STRING | EOX93440 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM7949 | 26 | 39 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G04880.2 | 5e-96 | zinc-dependent activator protein-1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG002318t1 |
| Entrez Gene | 18612431 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



